Imaging and structural studies of DNA–protein complexes and membrane ion channels†
Abstract
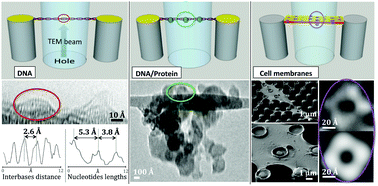
In bio-imaging by electron microscopy, damage of the sample and limited contrast are the two main hurdles for reaching high image quality. We extend a new preparation method based on nanofabrication and super-hydrophobicity to the imaging and structural studies of nucleic acids, nucleic acid–protein complexes (DNA/Rad51 repair protein complex) and neuronal ion channels (gap-junction, K+ and GABAA channels) as paradigms of biological significance and increasing complexity. The preparation method is based on the liquid phase and is compatible with physiological conditions. Only in the very last stage, samples are dried for TEM analysis. Conventional TEM and high-resolution TEM (HRTEM) were used to achieve a resolution of 3.3 and 1.5 Å, respectively. The EM dataset quality allows the determination of relevant structural and metrological information on the DNA structure, DNA–protein interactions and ion channels, allowing the identification of specific macromolecules and their structure.


 Please wait while we load your content...
Please wait while we load your content...