BiKEGG: a COBRA toolbox extension for bridging the BiGG and KEGG databases†
Abstract
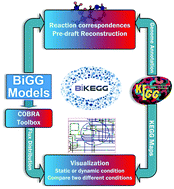
Development of an interface tool between the Biochemical, Genetic and Genomic (BiGG) and KEGG databases is necessary for simultaneous access to the features of both databases. For this purpose, we present the BiKEGG toolbox, an open source COBRA toolbox extension providing a set of functions to infer the reaction correspondences between the KEGG reaction identifiers and those in the BiGG knowledgebase using a combination of manual verification and computational methods. Inferred reaction correspondences using this approach are supported by evidence from the literature, which provides a higher number of reconciled reactions between these two databases compared to the MetaNetX and MetRxn databases. This set of equivalent reactions is then used to automatically superimpose the predicted fluxes using COBRA methods on classical KEGG pathway maps or to create a customized metabolic map based on the KEGG global metabolic pathway, and to find the corresponding reactions in BiGG based on the genome annotation of an organism in the KEGG database. Customized metabolic maps can be created for a set of pathways of interest, for the whole KEGG global map or exclusively for all pathways for which there exists at least one flux carrying reaction. This flexibility in visualization enables BiKEGG to indicate reaction directionality as well as to visualize the reaction fluxes for different static or dynamic conditions in an animated manner. BiKEGG allows the user to export (1) the output visualized metabolic maps to various standard image formats or save them as a video or animated GIF file, and (2) the equivalent reactions for an organism as an Excel spreadsheet.
 Please wait while we load your content...
Please wait while we load your content...