Invader probes: harnessing the energy of intercalation to facilitate recognition of chromosomal DNA for diagnostic applications†
Abstract
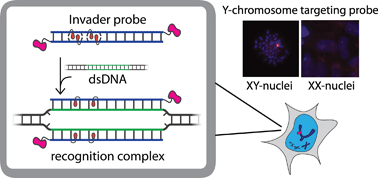
Development of probes capable of recognizing specific regions of chromosomal DNA has been a long-standing goal for chemical biologists. Current strategies such as PNA, triplex-forming oligonucleotides, and polyamides are subject to target choice limitations and/or necessitate non-physiological conditions, leaving a need for alternative approaches. Toward this end, we have recently introduced double-stranded oligonucleotide probes that are energetically activated for DNA recognition through modification with +1 interstrand zippers of intercalator-functionalized nucleotide monomers. Herein, probes with different chemistries and architectures – varying in the position, number, and distance between the intercalator zippers – are studied with respect to hybridization energetics and DNA-targeting properties. Experiments with model DNA targets demonstrate that optimized probes enable efficient (C50 < 1 μM), fast (t50 < 3 h), kinetically stable (>24 h), and single nucleotide specific recognition of DNA targets at physiologically relevant ionic strengths. Optimized probes were used in non-denaturing fluorescence in situ hybridization experiments for detection of gender-specific mixed-sequence chromosomal DNA target regions. These probes present themselves as a promising strategy for recognition of chromosomal DNA, which will enable development of new tools for applications in molecular biology, genomic engineering and nanotechnology.

 Please wait while we load your content...
Please wait while we load your content...