Preliminary investigation of metabolic engineering in a novel host bacterium Corynebacterium crenatum for alcohol biofuel production
Abstract
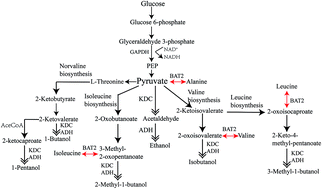
Bio-engineered microorganisms have been proposed as candidates to produce alcohols that can replace dwindling fossil fuel resources. In this paper, we are the first to investigate using a novel engineered bacterium, Corynebacterium crenatum, as a potential cell factory to produce alcohols. We investigated the metabolic engineering of Corynebacterium crenatum by testing the heterologous expression of 2-keto acid decarboxylases and branched-chain amino transferases. We verified that C. crenatum can produce alcohol biofuels by over-expressing several tandem KDCs (2-keto acid decarboxylases) and alcohol dehydrogenases derived from different yeast species. In addition, BAT2 gene (branched-chain amino acid transaminase) from Saccharomyces cerevisiae have also been introduced and investigated the effect of producing alcohols. We successfully enabled this C. crenatum strain to produce ethanol, including the higher alcohols isobutanol, butanol, and 3-methyl-1-butanol. Our results demonstrate that when the kivd gene from Lactococcus lactis subsp. cremoris and the BAT2 gene from S. cerevisiae are simultaneously over-expressed in C. crenatum, ethanol can be produced at concentrations of 5.85 g L−1 from hexose glucose, 7.28 g L−1 from pentose xylose and 6.38 g L−1 from mixed sugars. Additional alcohols can be produced at the following concentrations: isobutanol at 193.66 mg L−1, butanol at 173.07 mg L−1, 3-methyl-1-butanol at 449.60 mg L−1, and pentanol at 28.61 mg L−1. Our results demonstrate that this novel bioengineered strain of C. crenatum can produce a mixture of alcohols and seems to preferentially produce ethanol. Our results also suggest that it may be possible to use this bacterium to synthesize higher alcohols. Our preliminary investigation demonstrates it is possible to develop new bacterial hosts as metabolic engineering platforms for the production of biofuel energy.

 Please wait while we load your content...
Please wait while we load your content...