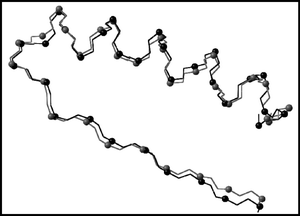
We apply a computational scheme, in which geometrical (through-space) and topological (through-bond) distances are combined to construct geometry-dependent structural D/D matrices, in order to develop a matrix invariant characterization of Top7, a de novo sequence design by Kuhlman et al. (B. Kuhlman, G. Dantas, G. C. Ireton. G. Varani, B. L. Stoddard and D. Baker, Science, 2003, 302, 1364), which was shown to exhibit a novel α/β globular protein fold. A sequence of invariants KΦ, known as the “folding profile” of a protein, are extracted from the D/D matrix and its related higher order representations, and a similarity measure is introduced allowing invariant-based quantification of similarity between protein forms. Quantitative comparison of the Top7 crystal structure with the computationally designed model is achieved for the full protein backbone and a specific substructural segment: Lys46–Tyr76. The approach appears particularly attractive as it allows both global comparisons, as well as characterization of local structural features of proteins.
You have access to this article
 Please wait while we load your content...
Something went wrong. Try again?
Please wait while we load your content...
Something went wrong. Try again?


 Please wait while we load your content...
Please wait while we load your content...