Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Real-time monitoring of photoinduced atom transfer radical polymerization by time-resolved diffusion NMR
Marek
Czarnota
 a,
Martyna
Cybularczyk-Cecotka
a,
Martyna
Cybularczyk-Cecotka
 b,
Franz F.
Westermair
b,
Franz F.
Westermair
 c,
Francisco
Arrabal Campos
c,
Francisco
Arrabal Campos
 d,
Kriti
Kapil
d,
Kriti
Kapil
 e,
Krzysztof
Matyjaszewski
e,
Krzysztof
Matyjaszewski
 e,
Grzegorz
Szczepaniak
e,
Grzegorz
Szczepaniak
 *b and
Mateusz
Urbańczyk
*b and
Mateusz
Urbańczyk
 *a
*a
aInstitute of Physical Chemistry, Polish Academy of Sciences, Kasprzaka 44/52, 01-224 Warszawa, Poland. E-mail: murbanczyk@ichf.edu.pl
bFaculty of Chemistry, University of Warsaw, Pasteura 1, 02-093 Warszawa, Poland. E-mail: g.szczepaniak@uw.edu.pl
cFaculty of Chemistry and Pharmacy, University of Regensburg, Universitätsstraße 31, 93053 Regensburg, Germany
dDepartment of Engineering, Research Centre CIAIMBITAL, University of Almería, 04120 Almería, Spain
eDepartment of Chemistry, Carnegie Mellon University, Pittsburgh, Pennsylvania 15213, USA
First published on 18th February 2026
Abstract
Conventional real-time monitoring of polymerization is a laborious process that requires offline sampling and analysis. In this work, we introduce an advanced approach that combines Time-Resolved Diffusion NMR with 1H NMR for the simultaneous, in situ tracking of both monomer conversion and molecular weight evolution. We successfully applied this method to monitor the photoinduced atom transfer radical polymerization (photo-ATRP) in real-time. Furthermore, by coupling the gamma model with dispersity-matched accumulation, we accurately determined the dispersity values of the resulting polymers. To make the method more accessible, we developed a user-friendly program that sets up the acquisition and simplifies real-time monitoring of photo-ATRP. Additionally, the program allows stopping illumination at any desired monomer conversion, enabling easy control of the process.
Introduction
Synthetic polymers represent one of the most widely used classes of chemical compounds, enabling applications from everyday packaging to advanced drug delivery systems. The precise control over their properties, such as molecular weight, architecture, and functionality, is paramount. This necessity has driven the development of reversible-deactivation radical polymerization (RDRP) techniques,1,2 with atom transfer radical polymerization (ATRP) standing out as a particularly powerful and versatile method since its discovery in 1995.3,4However, conventional ATRP requires high concentrations of an air-sensitive copper(I) catalyst. To overcome this limitation, photoinduced ATRP (photo-ATRP) was developed,5 which leverages light as a clean and efficient external stimulus to mediate the polymerization. This approach provides exceptional spatiotemporal control,6 as the reaction can be initiated and halted simply by switching a light source on or off. Furthermore, photo-ATRP can be conducted in green solvents,7 including water,8 broadening the scope of compatible monomers and initiators.9,10 It also exhibits greater tolerance to molecular oxygen,11 eliminating the need for stringent deoxygenation procedures.
These features make photo-ATRP a highly promising method for both academic research and industrial manufacturing. However, its implementation and scalability are hindered by a critical challenge: photochemical reactions are extremely sensitive to subtle variations in experimental conditions.12 Factors like light source intensity, reactor geometry, distance from the light source, and the homogeneity of the reaction mixture can profoundly impact polymerization kinetics and its reproducibility.13 Minor, often unavoidable, deviations between experiments can lead to inconsistent outcomes, creating a significant barrier to the use of this technique.
This inherent sensitivity underscores a pressing need for robust analytical methods capable of monitoring the reaction's progress in real-time and under true process conditions (in situ).14 Such tools are essential for understanding reaction kinetics, ensuring reproducibility, and enabling the effective optimization of polymerization conditions. The ongoing quest to develop tools for online polymerization monitoring has led to several key methods,15 including electrospray ionization mass spectrometry,16infra-red spectroscopy,17 chromatography,18 and nuclear magnetic resonance (NMR).19
NMR monitoring has become one of the workhorses of modern chemists. However, this field is still growing in terms of methodology, showing progress from basic 1H one-dimensional series of spectra, e.g., ref. 20 and 21 to different approaches like multidimensional spectroscopy for more complex systems.22–28
In the case of polymerization, researchers usually utilize 1H NMR to monitor monomer conversion.19,29,30 Additional approaches which can be used are 13C MAS NMR,31 or the Diffusion-NMR which is rapidly being utilized to monitor time-dependent processes.27,28,32 This class of NMR is based on the analysis of the diffusion coefficient change in time instead of conventional frequency-based analysis. This approach allows a better insight into a process where the peak structure does not change (or changes insignificantly) while the molecular weight or freedom of movement is evolving. One of the most popular classes of reactions that fit the system above is polymerization. Usually, the molecular weight of the polymer changes significantly while the NMR spectrum of the polymer stays the same.
For such a system, diffusion NMR seems to be an ideal solution as the diffusion coefficient is linked to the molecular weight.33 However, conventional diffusion NMR is not best suited for reaction monitoring. As the measurement time of a single experiment varies from minutes to hours, the temporal information can be lost. Additionally, the change in the concentration of reagents can affect the measured diffusion coefficients.31
Two acquisition strategies have been implemented to overcome this problem. The first, based on the spatial encoding of the diffusion coefficient, is called ultra-fast.28,34,35 The second is based on the time-resolved non-uniform sampling,36 where instead of randomly sampling the t1 evolution time, one samples pulsed field gradients in the PGSTE sequence.37,38 Both methods have strengths and weaknesses, and choosing the proper acquisition strategy is essential for the proper analysis.39
In this work, we utilized the Time-Resolved Diffusion NMR (TR-DNMR) as this method has proven successful for monitoring polymerization processes.27,40 Additionally, the photo-ATRP process is light-driven; therefore, to study the process in situ, illumination must be provided to the NMR tube. The NMR community has proposed several approaches.41–44 Currently, the most commonly used solution is to provide the illumination using an optical fiber with a polished tip that is inserted into the NMR tube.45 This allows for uniform illumination of the sample volume and enables easy switching of the light source.
Results and discussion
In situ polymerization and light control
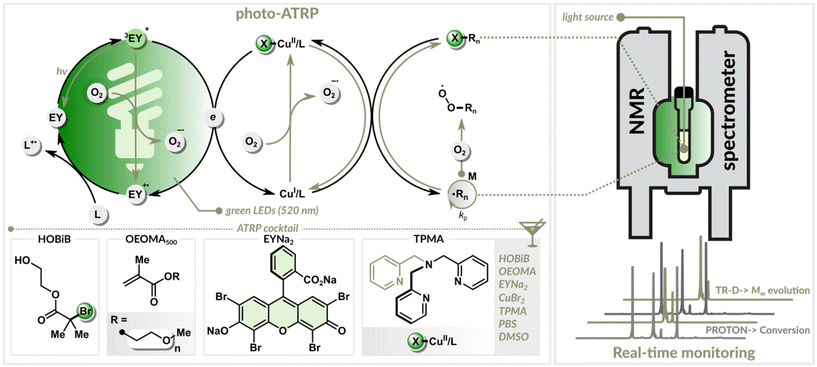
We employed a photo-ATRP method that combines organic photoredox catalysis with copper catalysis (Fig. 1).30,46 A key advantage of this dual catalytic system is its high oxygen tolerance, which enables efficient polymerization under green light illumination without needing prior deoxygenation.30 This approach yields well-defined polymers over a wide range of targeted degrees of polymerization (DPT).47–49To monitor the reaction in real-time, polymerizations were conducted inside an NMR tube illuminated by a centrally positioned optical fiber. A portion of the fiber was stripped from its cladding and polished. A New Era Photo-NMR device was used to ensure a stable fiber alignment, minimizing interference with magnetic field homogeneity. The polished end of the fiber was inserted in a way that the whole sample was evenly illuminated. The LED illumination (520 nm) was controlled by an Arduino module, while a custom-written program with a graphical user interface (GUI) managed the experiment. This automated system provided real-time monitoring of monomer conversion, polymer diffusion coefficient, and molecular weight evolution. The system was programmed to automatically halt the polymerization by turning off the light once a predefined monomer conversion was achieved.
To demonstrate the effectiveness of this in situ monitoring technique, we performed a series of polymerizations (Table 1). The experiments used oligo(ethylene glycol) methyl ether methacrylate (OEOMA500) as the benchmark water-soluble monomer. The other components were 2-hydroxyethyl 2-bromoisobutyrate (HOBiB) as the initiator, eosin Y disodium salt (EYNa2) as the organic photoredox catalyst, and a CuBr2/TPMA complex (TPMA = tris(2-pyridylmethyl)amine) as the copper catalyst. The polymerizations were conducted in phosphate-buffered saline (PBS 1X) containing 10% v/v non-deuterated dimethyl sulfoxide (DMSO). The targeted DP was varied by adjusting the HOBiB concentration (6.0–0.375 mM), while all other component concentrations remained fixed (Table 1).
| Entry | DPT | Conv.![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) b b |
M
n,th![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) c c |
M w,SEC | M n,SEC | Đ SEC |
Đ
NMR![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) f f |
|---|---|---|---|---|---|---|---|
| a Reaction conditions [OEOMA500] = 300 mM, [HOBiB] = 6.0–0.375 mM, [EYNa2] = 15 μM, [CuBr2] = 0.3 mM, [TPMA] = 0.9 mM in PBS with DMSO (10% v/v), irradiated under green LEDs (520 nm) in NMR tube. b Monomer conversion was determined by using 1H NMR spectroscopy. c M n,th = DPT × conv. × MWOEOMA500 + MWHOBIB. d Molecular weight (Mn,SEC and Mw,SEC) and dispersity (ĐSEC) were determined by size-exclusion chromatography (SEC) with a multi-angle light scattering (MALS) detector (DMF as eluent). e M w,SEC, Mn,SEC and ĐSEC were determined by Mark–Houwink calibration. f Dispersity (ĐNMR) was determined by NMR diffusion measurements. | |||||||
| 1d | 50 | 90% | 22![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 700 700 |
26![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 400 400 |
21![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 900 900 |
1.20 | 1.17 |
| 2d | 100 | 90% | 45![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 200 200 |
56![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 000 |
45![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 300 300 |
1.24 | 1.25 |
| 3d | 200 | 90% | 90![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 200 200 |
96![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 300 300 |
84![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 500 500 |
1.14 | 1.08 |
| 4e | 400 | 70% | 140![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 200 200 |
158![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 700 700 |
139![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 100 100 |
1.11 | 1.12 |
| 5e | 800 | 70% | 280![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 200 200 |
374![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 900 900 |
251![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 100 100 |
1.39 | 1.37 |
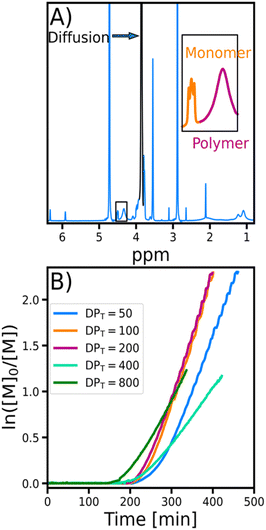
The EY/Cu-catalyzed photo-ATRP process began with an induction period of approximately 200 min, during which dissolved oxygen was consumed (Fig. 2B). In this initial stage, oxygen quenched the excited photocatalyst and oxidized the copper(I) activator (CuI/L) (Fig. 1). Once the oxygen concentration dropped below a critical threshold, the polymerization started. The excited photocatalyst reduced the copper(II) deactivator (X–CuII/L), regenerating the CuI/L activator needed for polymer growth. Precise control was maintained through the reversible equilibrium between the activating Cu(I) and deactivating Cu(II) species, ensuring controlled, uniform growth of polymer chains (Fig. 1).50
Achieving precise control over monomer conversion is crucial for tailoring a polymer's final physical properties, such as its viscosity, glass transition temperature, and mechanical behavior. We demonstrated the versatility of our in situ NMR monitoring method by automatically stopping the light illumination once the desired monomer conversion was achieved. For reactions targeting a DP between 50 and 200, the process was stopped at 90% monomer conversion. For higher DPT of 400 and 800, the reactions were halted at 70% conversion to diminish the dead-chain fraction (defined as the ratio of the concentration of terminated polymer chains to the initial concentration of the initiator).51 As shown in Table 1, the resulting polymers exhibited low dispersity values and molecular weights (Mn,SEC) in close agreement with theoretical values (Mn,th), confirming the well-controlled nature of this technique.
Time-resolved diffusion NMR
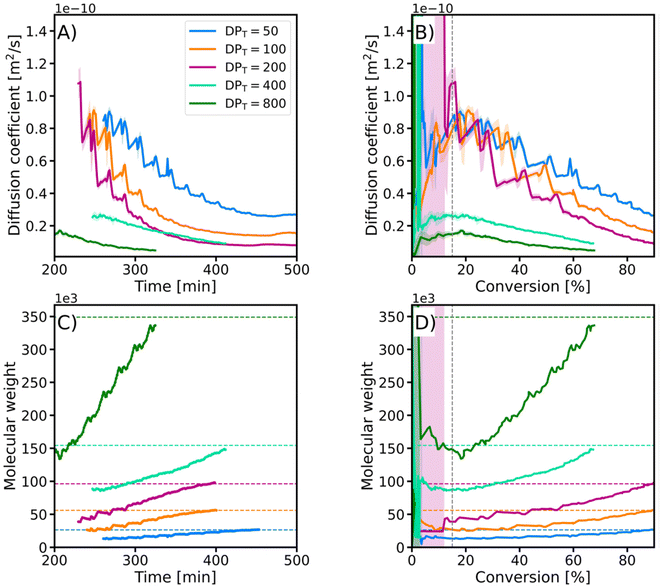
To monitor the evolution of molecular weight during the polymerization (Fig. 2), Time-Resolved Diffusion NMR (TR-DNMR) was used.37,38 This method combines the diffusion NMR equivalent of the Time-Resolved Non-Uniform Sampling (TR-NUS)36,52,53 and permuted Diffusion Ordered Spectroscopy (p-DOSY) concepts.54The p-DOSY technique was introduced to eliminate the interference from the kinetics on the diffusion coefficient. The concept is simple: instead of sampling the pulsed field gradient strengths in ascending order, they are permuted. This makes the signal intensity change due to kinetics independent of the gradient strength. The TR-NUS technique was introduced to enable the monitoring of reactions with 2D NMR. In this approach, the evolution step of the indirect dimension is randomly sampled during the reaction. After the acquisition, the data are artificially separated into overlapping subsets, and the whole spectra are reconstructed.55 Because the average time interval between these subsets is very short, the reconstructed data allow the reaction to be followed with high temporal resolution. We combined both concepts to monitor the evolution of the diffusion coefficient during the polymerization with excellent temporal resolution (the interval between points was 1 m 12 s).
In TR-DNMR, the concept is based on a series of PGSTE experiments, where the series of gradient strengths is permuted and repeated throughout the reaction time. Then, the data are divided into p-DOSY subsets, which are used to calculate the diffusion coefficients by fitting a mono-exponential model or an inverse Laplace transform.56 Additionally, this approach allows the use of interleaved (IL) acquisition,25 in which each step of the TR-DNMR experiment can be combined with a different one. In this work, we have interleaved the diffusion experiments with 1D proton experiments to calculate the monomer conversions.
The diffusion coefficients (D) were derived from TR-DNMR experiments. The experiments were divided into subsets of size 16, overlapping with each other by 15 experiments, so that each subset contained the same set of gradients. To determine the polymer diffusion coefficient (Dp), the peak at 3.85 ppm was integrated and its intensity (I(b)) was fitted with a double exponential model:
| I(b) = I0Ce−Dp×b + I0(1 − C)e−Dm×b | (1) |
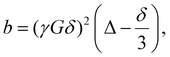 | (2) |
In the early stages of the polymerization (conv. <15%), the diffusion coefficient estimation had a significant fitting error due to the low intensity of the polymer signal compared to that of the monomer. However, after this initial disturbance, the evolution of the diffusion coefficient followed a similar path for all samples. The regular small fluctuations of the diffusion coefficients were caused by the repetitive sampling of pulsed field gradients. Additionally, we calculated the evolution of the weight-average molecular weight (Mw) from the diffusion coefficients using the equation:
| D = kMwα, | (3) |
Dispersity-matched accumulation diffusion NMR
NMR diffusion measurements can be used to calculate the polymer dispersity (Đ).37,62,63 This method relies on modelling the non-monoexponential decay of the diffusion signal, for which a gamma distribution model can be used.62 The data are fitted using a modified version of the conventional Stejskal-Tanner equation:64| I(b) = I0(1 + bσ2/D)(−D2/σ2) | (4) |
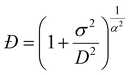 | (5) |
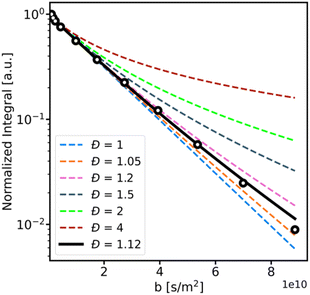
However, as shown in Fig. 4, for samples with dispersity values below 2, the deviation from the classical mono-exponential model is visible only for high gradients, where the signal-to-noise ratio is significantly diminished. This causes difficulty in fitting the gamma model for experiments with a moderate number of scans and necessitates extremely long experiments with many transient acquisitions. Such an approach is far from optimal, especially for the points with low gradient strength, where the SNR is high, and multiple scans do not increase the data quality.
To overcome this issue, we propose an adaptation of the matched accumulation concept for diffusion dispersity measurements.65 In this method, the number of scans is adapted for each gradient strength to maintain a constant SNR across the measurement. We analyzed the dispersity of the post-reaction samples using this combined approach of a gamma-model distribution and dispersity-matched accumulation. The resulting values were compared to those determined by SEC analysis (see Table 1) and showed excellent agreement.
Conclusions
In conclusion, we demonstrated the efficacy of the Time-Resolved Diffusion NMR method for monitoring the photo-ATRP in real time. Our approach allowed us to track the evolution of the diffusion coefficient and molecular weight of the polymer as the reaction proceeded under illumination. To facilitate wider adoption of this method, we developed a user-friendly program that automated reaction monitoring and enabled the polymerization to be halted at a precise monomer conversion. Finally, we showed that the Diffusion NMR, when combined with a gamma-model and dispersity-matched accumulation, is a powerful alternative to conventional size exclusion chromatography (SEC) measurements, allowing the study of the process in situ with significantly less effort from the operator. Additionally, the method might be combined with a benchtop NMR spectrometer and flow setup to monitor the process inside the full-scale reactor for larger-scale polymerization.Author contributions
M. C.: methodology, software, formal analysis, investigation, visualization, writing – original draft. M. C. C.: investigation, formal analysis, writing – original draft, writing – review and editing. F. F. W.: software. F. A. C.: investigation. K. K.: investigation, formal analysis. K. M.: writing – review and editing. G. S.: conceptualization, methodology, resources, writing – original draft, writing – review and editing, supervision, project administration, funding acquisition. M. U.: conceptualization, methodology, resources, writing – original draft, writing – review and editing, supervision, project administration, funding acquisition.Conflicts of interest
There are no conflicts to declare.Data availability
The NMR raw data and processing scripts used for the results shown in this manuscript are available in RepOD ICM repository at: https://doi.org/10.18150/RB78J7. The acquisition and live monitoring programs are available on GitHub at: https://github.com/murbanczyk/PhotopolymerizationMonitor.Supplementary information (SI) is available. See DOI: https://doi.org/10.1039/d6py00103c.
Acknowledgements
M. U. and M. C would like to thank the National Science Centre, Poland, for its support in the form of an OPUS Grant (2021/41/B/ST4/01286). M. C. C and G. S acknowledge support from the Polish National Agency for Academic Exchange under the Polish Returns program (BPN/PPO/2022/1/00027) and the Polish National Science Center (2023/02/1/ST4/00009). K. K. and K. M. acknowledge support from NSF (CHE 2401112).References
- N. Corrigan, K. Jung, G. Moad, C. J. Hawker, K. Matyjaszewski and C. Boyer, Prog. Polym. Sci., 2020, 111, 101311 CrossRef CAS.
- K. Parkatzidis, H. S. Wang, N. P. Truong and A. Anastasaki, Chem, 2020, 6, 1575–1588 CAS.
- J. S. Wang and K. Matyjaszewski, J. Am. Chem. Soc., 1995, 117, 5614–5615 CrossRef CAS.
- S. Harrisson, R. Whitfield, A. Anastasaki and K. Matyjaszewski, Nat. Rev. Methods Primers, 2025, 5, 1–20 Search PubMed.
- X. Pan, M. A. Tasdelen, J. Laun, T. Junkers, Y. Yagci and K. Matyjaszewski, Prog. Polym. Sci., 2016, 62, 73–125 CrossRef CAS.
- S. Dadashi-Silab, I. H. Lee, A. Anastasaki, F. Lorandi, B. Narupai, N. D. Dolinski, M. L. Allegrezza, M. Fantin, D. Konkolewicz, C. J. Hawker and K. Matyjaszewski, Macromolecules, 2020, 53, 5280–5288 CrossRef CAS.
- S. Dworakowska, F. Lorandi, A. Gorczyński and K. Matyjaszewski, Adv. Sci., 2022, 9, 2106076 Search PubMed.
- X. Pan, N. Malhotra, A. Simakova, Z. Wang, D. Konkolewicz and K. Matyjaszewski, J. Am. Chem. Soc., 2015, 137, 15430–15433 Search PubMed.
- R. A. Olson, A. B. Korpusik and B. S. Sumerlin, Chem. Sci., 2020, 11, 5142–5156 Search PubMed.
- F. Lorandi, M. Fantin, H. Jafari, A. Gorczyński, G. Szczepaniak, S. Dadashi-Silab, A. A. Isse and K. Matyjaszewski, J. Am. Chem. Soc., 2023, 145, 21587–21599 CrossRef CAS PubMed.
- G. Szczepaniak, L. Fu, H. Jafari, K. Kapil and K. Matyjaszewski, Acc. Chem. Res., 2021, 54, 1779–1790 CrossRef CAS PubMed.
- M. A. Beres, C. Boyer, M. Hartlieb, D. Konkolewicz, G. G. Qiao, B. S. Sumerlin and S. Perrier, ACS Polym. Au, 2025, 5, 184–213 CrossRef CAS PubMed.
- T. D. Svejstrup, A. Chatterjee, D. Schekin, T. Wagner, J. Zach, M. J. Johansson, G. Bergonzini and B. König, ChemPhotoChem, 2021, 5, 808–814 CrossRef CAS.
- O. Tooley, W. Pointer, R. Radmall, M. Hall, T. Swift, J. Town, C. Aydogan, T. Junkers, P. Wilson, D. Lester and D. Haddleton, ACS Polym. Au, 2024, 4, 311–319 Search PubMed.
- J. J. Haven and T. Junkers, Eur. J. Org. Chem., 2017, 2017, 6474–6482 CrossRef CAS.
- J. J. Haven, J. Vandenbergh and T. Junkers, Chem. Commun., 2015, 51, 4611–4614 RSC.
- J. E. Puskas, P. Chan, K. B. McAuley, G. Kaszas and S. Shaikh, Macromol. Symp., 2006, 240, 18–22 CrossRef CAS.
- R. Hoogenboom, M. W. M. Fijten, C. H. Abeln and U. S. Schubert, Macromol. Rapid Commun., 2004, 25, 237–242 CrossRef CAS.
- M. Rubens, J. Van Herck and T. Junkers, ACS Macro Lett., 2019, 8, 1437–1441 Search PubMed.
- X. L. Wang, W. Liu, Y. Y. Yu, Y. Song, W. Q. Fang, D. Wei, X. Q. Gong, Y. F. Yao and H. G. Yang, Nat. Commun., 2016, 7, 1–7 Search PubMed.
- D. A. Foley, E. Bez, A. Codina, K. L. Colson, M. Fey, R. Krull, D. Piroli, M. T. Zell and B. L. Marquez, Anal. Chem., 2014, 86, 12008–12013 Search PubMed.
- R. Dass, W. Koźmiński and K. Kazimierczuk, Anal. Chem., 2015, 87, 1337–1343 Search PubMed.
- C. Jacquemmoz, F. Giraud and J.-N. Dumez, Analyst, 2020, 145, 478–485 RSC.
- B. Gouilleux, B. Charrier, E. Danieli, J. N. Dumez, S. Akoka, F. X. Felpin, M. Rodriguez-Zubiri and P. Giraudeau, Analyst, 2015, 140, 7854–7858 Search PubMed.
- M. Urbańczyk, A. Shchukina, D. Gołowicz and K. Kazimierczuk, Magn. Reson. Chem., 2019, 57, 4–12 Search PubMed.
- D. Gołowicz, K. Kazimierczuk, M. Urbańczyk and T. Ratajczyk, ChemistryOpen, 2019, 8, 196–200 Search PubMed.
- K. Kristinaityte, A. Mames, M. Pietrzak, F. F. Westermair, W. Silva, R. M. Gschwind, T. Ratajczyk and M. Urbańczyk, J. Am. Chem. Soc., 2022, 144, 13938–13945 CrossRef CAS PubMed.
- V. V. Telkki, M. Urbańczyk and V. Zhivonitko, Prog. Nucl. Magn. Reson. Spectrosc., 2021, 126–127, 101–120 CrossRef CAS PubMed.
- L. L. Jessen, K. R. Hansen, G. B. Crull, T. L. Grover and C. A. Guymon, ACS Macro Lett., 2025, 14, 847–852 CrossRef CAS PubMed.
- G. Szczepaniak, J. Jeong, K. Kapil, S. Dadashi-Silab, S. S. Yerneni, P. Ratajczyk, S. Lathwal, D. J. Schild, S. R. Das and K. Matyjaszewski, Chem. Sci., 2022, 13, 11540–11550 RSC.
- A. Lasorsa, P. van der Meulen, E. Naumann, M. M. Lerch, R. M. Garcia, X. Lan, K. Loos, B. L. Feringa, W. Szymanski and P. C. A. van der Wel, J. Mater. Chem. A, 2025, 13, 36318–36329 RSC.
- S. Ahola and V.-V. Telkki, ChemPhysChem, 2014, 15, 1687–1692 Search PubMed.
- O. Tooley, W. Pointer, R. Radmall, M. Hall, V. Beyer, K. Stakem, T. Swift, J. Town, T. Junkers, P. Wilson, D. Lester and D. Haddleton, Macromol. Rapid Commun., 2024, 45, 2300692 CrossRef CAS PubMed.
- L. Guduff, I. Kuprov, C. Van Heijenoort and J. N. Dumez, Chem. Commun., 2017, 53, 701–704 Search PubMed.
- G. Hamdoun, L. Guduff, C. Van Heijenoort, C. Bour, V. Gandon and J. N. Dumez, Analyst, 2018, 143, 3458–3464 RSC.
- M. Mayzel, J. Rosenlöw, L. Isaksson and V. Y. Orekhov, J. Biomol. NMR, 2014, 58, 129–139 CrossRef CAS PubMed.
- M. Urbańczyk, D. Bernin, A. Czuroń and K. Kazimierczuk, Analyst, 2016, 141, 1745–1752 RSC.
- T. S. C. MacDonald, W. S. Price and J. E. Beves, ChemPhysChem, 2019, 20, 926–930 CrossRef CAS PubMed.
- M. Urbańczyk, Y. Kharbanda, O. Mankinen and V.-V. Telkki, Anal. Chem., 2020, 92, 9948–9955 CrossRef PubMed.
- L. L. Fillbrook, M. D. Nothling, M. H. Stenzel, W. S. Price and J. E. Beves, ACS Macro Lett., 2022, 11, 166–172 CrossRef CAS PubMed.
- J. E. Bramham and A. P. Golovanov, Commun. Chem., 2022, 5, 1–10 Search PubMed.
- P. Nitschke, N. Lokesh and R. M. Gschwind, Prog. Nucl. Magn. Reson. Spectrosc., 2019, 114–115, 86–134 CrossRef CAS PubMed.
- A. Seegerer, P. Nitschke and R. M. Gschwind, Angew. Chem., Int. Ed., 2018, 57, 7493–7497 CrossRef CAS PubMed.
- Y. Ji, D. A. DiRocco, J. Kind, C. M. Thiele, R. M. Gschwind and M. Reibarkh, ChemPhotoChem, 2019, 3, 984–992 CrossRef CAS.
- C. Feldmeier, H. Bartling, E. Riedle and R. M. Gschwind, J. Magn. Reson., 2013, 232, 39–44 CrossRef CAS PubMed.
- J. Sobieski, A. Gorczyński, A. M. Jazani, G. Yilmaz and K. Matyjaszewski, Angew. Chem., Int. Ed., 2025, 64, e202415785 CrossRef CAS PubMed.
- R. Bernat, G. Szczepaniak, K. Kamiński, M. Paluch, K. Matyjaszewski and P. Maksym, Chem. Commun., 2024, 60, 843–846 RSC.
- K. Kapil, G. Szczepaniak, M. R. Martinez, H. Murata, A. M. Jazani, J. Jeong, S. R. Das and K. Matyjaszewski, Angew. Chem., Int. Ed., 2023, 62, e202217658 Search PubMed.
- J. Jeong, G. Szczepaniak, S. R. Das and K. Matyjaszewski, Chem, 2023, 9, 3319–3334 CAS.
- F. Lorandi, M. Fantin and K. Matyjaszewski, J. Am. Chem. Soc., 2022, 144, 15413–15430 CrossRef CAS PubMed.
- K. Matyjaszewski, Macromolecules, 2012, 45, 4015–4039 CrossRef CAS.
- D. Gołowicz, P. Kasprzak, V. Orekhov and K. Kazimierczuk, Prog. Nucl. Magn. Reson. Spectrosc., 2020, 116, 40–55 Search PubMed.
- A. Shchukina, M. Urbańczyk, P. Kasprzak and K. Kazimierczuk, Concepts Magn. Reson., Part A: Bridging Educ. Res., 2017, 46, e21429 CrossRef.
- M. Oikonomou, J. Asencio-Hernández, A. H. Velders and M.-A. Delsuc, J. Magn. Reson., 2015, 258, 12–16 CrossRef CAS PubMed.
- K. Kazimierczuk and V. Y. Orekhov, Angew. Chem., Int. Ed., 2011, 50, 5556–5559 CrossRef CAS PubMed.
- M. Urbańczyk, D. Bernin, W. Koźmiński and K. Kazimierczuk, Anal. Chem., 2013, 85, 1828–1833 CrossRef PubMed.
- W. Li, H. Chung, C. Daeffler, J. A. Johnson and R. H. Grubbs, Macromolecules, 2012, 45, 9595–9603 CrossRef CAS PubMed.
- B. Grabe and W. Hiller, Macromolecules, 2022, 55, 8014–8020 CrossRef CAS.
- W. Hiller, B. Grabe and J. Schonert, Anal. Chem., 2024, 96, 14902–14908 Search PubMed.
- F. M. Arrabal-Campos, P. Oña-Burgos and I. Fernández, Polym. Chem., 2016, 7, 4326–4329 RSC.
- P. J. Voorter, A. McKay, J. Dai, O. Paravagna, N. R. Cameron and T. Junkers, Angew. Chem., Int. Ed., 2022, 61, e202114536 CrossRef CAS PubMed.
- M. Röding, D. Bernin, J. Jonasson, A. Särkkä, D. Topgaard, M. Rudemo and M. Nydén, J. Magn. Reson., 2012, 222, 105–111 CrossRef PubMed.
- P. Mocny, T. C. Lin, R. Parekh, Y. Zhao, M. Czarnota, M. Urbańczyk, C. Majidi and K. Matyjaszewski, ACS Appl. Mater. Interfaces, 2024, 16, 23932–23947 CAS.
- E. O. Stejskal and J. E. Tanner, J. Chem. Phys., 1965, 42, 288 CrossRef CAS.
- A. Kumar, S. C. Brown, M. E. Donlan, B. U. Meier and P. W. Jeffs, J. Magn. Reson., 1991, 95, 1–9 CAS.
| This journal is © The Royal Society of Chemistry 2026 |