Open Access Article
Open Access ArticleLipid adsorbed on the surface of nanomaterials enhances protein corona formation
Morgan M.
Heckman
 a,
Catherine S.
Cai
a,
Abhishek
Kalpattu
a,
Emily M.
Simmons
a,
Robert M.
Tighe
a,
Catherine S.
Cai
a,
Abhishek
Kalpattu
a,
Emily M.
Simmons
a,
Robert M.
Tighe
 b and
Christine K.
Payne
b and
Christine K.
Payne
 *a
*a
aThomas Lord Department of Mechanical Engineering and Materials Science, Duke University, Durham, North Carolina, USA 27708. E-mail: christine.payne@duke.edu
bDepartment of Medicine, Duke University School of Medicine, Durham, North Carolina, USA 27710
First published on 18th March 2026
Abstract
The high level of nanomaterials used in industrial and consumer products drives the need to better understand how these materials interact with biological systems. We probe the interaction of four commonly used nanomaterials; titanium dioxide, silicon dioxide, and silver nanoparticles and multi-walled carbon nanotubes, with a representative lung lining fluid lipid, L-α-phosphatidylcholine, and serum proteins. We find that the presence of the lipid increases the adsorption of fetal bovine serum on the surface of the nanomaterials. Individual serum proteins, bovine serum albumin and transferrin, show a nanomaterial-dependent response. We used murine macrophages to characterize the cellular response to lipid–protein–nanoparticle complexes and found a nanomaterial-dependent response, measured by cytokine release. In the case of titanium dioxide nanoparticles, L-α-phosphatidylcholine on the nanomaterial surface provides a protective effect against inflammation. While much recent work probes the protein corona that forms on nanomaterials used in biological applications, this work examines the lipid and protein coronas with relevance to inhalation exposures.
Environmental significanceNanomaterials are used in increasing quantities across a range of industries from construction to electronics to biomedical devices. This high level of use is associated with the risk of inhalation exposures, both for workers in a factory setting and the general public following environmental release or degradation of consumer products. We explore the interaction of lipids and proteins with four common nanomaterials (titanium dioxide, silicon dioxide, and silver nanoparticles and multi-walled carbon nanotubes) with relevance to inhalation exposures. Following inhalation, nanomaterials come into contact with both lipids and proteins in the lung lining fluid leading, to the formation of a lipid and protein corona. We examine the adsorption of lipids and proteins on the nanomaterial surface and the response of macrophages to the lipid–protein–nanomaterials. Overall, we hope this research will highlight the importance of lipids in nanomaterial exposures. |
Introduction
Nanomaterials are used at increasingly high levels in industrial and consumer products.1–7 This high level of use is associated with human exposure through inhalation in manufacturing settings and following environmental release.8–14 Of specific interest for human exposure are titanium dioxide nanoparticles (TiO2 NPs), used in plastics, coatings, cosmetics, sunscreen, and food products; silicon dioxide (SiO2) NPs used in optics, composites, coatings, and biomedical applications; silver (Ag) NPs used in antimicrobials and electronics; and multi-walled carbon nanotubes (MWCNTs) used in electronics and composites. In developed countries, most people will encounter these materials on a daily basis. In the United States, manufacturing exposure by inhalation has been regulated by the Occupational Safety and Health Administration (OSHA), which sets a permissible exposure limit (PEL) for workers over an 8 h workday.15 For consumer products containing these nanomaterials, the Food and Drug Administration (FDA) regulates exposure by dermal contact or ingestion. The Environmental Protection Agency (EPA) carries out case-by-case recommendations of regulations based on proposed use and potential environmental exposures. Common to all of these nanomaterials is the concern of inhalation exposure during production.16 OSHA regulates this for TiO2, SiO2, and Ag NPs. MWCNTs are considered respirable particles, but OSHA does not set a PEL.The first step in the inhalation of these nanomaterials is the interaction of the nanomaterial with lung lining fluid.17–25 Our lab,26,27 and others,28 have examined the interaction of lung lining fluid proteins with TiO2 NPs and MWCNTs. Other researchers have examined the interaction of SiO2 NPs with lung lining fluid.29 In general, this previous work has focused on protein–nanomaterial interactions and the formation of a “corona” of proteins on the nanomaterial surface. For example, we measured the concentration and composition of lung lining fluid proteins adsorbed on the surface of TiO2 NPs and found that the corona formed from lung lining fluid leads to elevated expression of pro-inflammatory cytokines.26
Less common are studies of the lipid corona; how lipids interact with nanomaterials, how they alter the protein interaction, and their importance for subsequent cellular responses. Consideration of lipid–nanomaterial interactions are especially important for inhalation exposures as lipids comprise 90% of the pulmonary surfactant in lung fluid.30,31 Previous work examining lipid–nanomaterial interactions has found that a lipid corona, including cholesterol and triglycerides, forms on polystyrene NPs (26 nm and 200 nm; anionic) incubated with mouse serum, which contains endogenous lipids.32 Incubation of gold NPs (14 nm) with lipid vesicles found an electrostatically-driven formation of a lipid corona.33 Early work in the field of lipid coronas relevant to inhalation exposures used porcine surfactant from lungs as a source of lipids. This work, using both proteomics and lipidomics, found that the lipid corona was independent of NP surface properties for polyethyleneglycol-iron oxide (160 nm), lipid (phosphatidylcholine)-iron oxide (169 nm) and PLGA-iron oxide NPs (177 nm). While the lipid composition was conserved, the protein coronas varied, especially proteins with lipid and surface binding properties.34 In a separate study, a combination of experimental and computational methods were used to probe the interaction of neutral, cationic, and anionic silica NPs (100 nm) with porcine lung surfactant, characterizing a lipid corona as a function of NP zeta potential, incubation time, and sonication.35
We describe a molecular and cellular study examining the interaction of lipids and proteins with TiO2, SiO2, Ag NPs and MWCNTs. We use L-α-phosphatidylcholine (LAP, lecithin) as a representative lipid, as we, and others,30,31 identified phosphatidylcholines as a major lipid type in human lung lining fluid. Serum proteins, including the mixture of proteins present in fetal bovine serum (FBS) and isolated albumin and transferrin, were used as a model protein system. Our experiments showed a surprising response: nanomaterials incubated with a mixture of LAP and blood serum proteins results in higher adsorption of proteins on the nanomaterial surface. The cellular response to the lipid–protein–nanomaterial complexes was measured by enzyme-linked immunosorbent assay (ELISA) using the generation of the common pro-inflammatory cytokine, tumor necrosis factor-α (TNF-α), as an indicator of a negative cellular response in macrophages. We find that LAP on the surface of nanoparticles leads to a nanomaterial-dependent response. In the case of TiO2 NPs, LAP on the surface of the NPs provides a protective effect for the cells. This research highlights the importance of controlled molecular studies to understand complex cellular responses to nanomaterials. We hope this work will inform regulations for human exposures and aid in the design of new, safer, nanomaterials.
Materials and methods
Nanomaterial characterization
Five different nanomaterials were used for experiments: carboxylate-modified magnetic NPs (magNPs; SC0200, Ocean NanoTech, San Diego, CA), TiO2 NPs (R101, 816719, Chemours, Fayetteville, NC), SiO2 NPs (637246-50G, MilliporeSigma, St. Louis, MO), Ag NPs (576832, MilliporeSigma), MWCNTs (NC7000, Nanocyl, Sambreville, Belgium). Nanomaterial characterization was carried out as previously described.36 Briefly, dynamic light scattering (DLS; Zetasizer, Malvern Instruments, Worcestershire, England) was used to measure the hydrodynamic diameter and polydispersity index (10 μg mL−1 in water following sonication; 5 min; Q700, Qsonica, Newtown, CT) and ζ-potential (10 μg mL−1 in 0.01x phosphate-buffered saline (PBS) in water) for all approximately spherical NPs (magNP, TiO2, SiO2, Ag NPs). Water is used for hydrodynamic diameter measurements to model industrial and environmental exposures. The 0.01x PBS salt solution provides a low ion strength solution for ζ-potential measurements.37 As DLS is not suitable for non-spherical particles, values for the MWCNTs are not included. Measurements were carried out in triplicate with three distinct samples. Average and standard deviations are reported for all measurements. Electrophoretic mobility was converted to ζ-potential using the Smoluchowski approximation. Primary particle diameters were measured from transmission electron microscopy (TEM; Tecnai G2 Twin; FEI; Hillsboro, Oregon; 25× to 700![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000× magnitude, 20 to 200 kV). Nanomaterials (1 mg mL−1 in water, 5 min sonication) were drop-cast onto carbon grids and dried overnight. TEM was carried out at Duke University's Shared Materials Instrumentation Facility. Primary particle diameters were measured for isolated particles that were not part of larger aggregates or from clearly visible boundaries within aggregates. A minimum of 50 particles from at least 3 images were analyzed. Image analysis was carried out with ImageJ.38
000× magnitude, 20 to 200 kV). Nanomaterials (1 mg mL−1 in water, 5 min sonication) were drop-cast onto carbon grids and dried overnight. TEM was carried out at Duke University's Shared Materials Instrumentation Facility. Primary particle diameters were measured for isolated particles that were not part of larger aggregates or from clearly visible boundaries within aggregates. A minimum of 50 particles from at least 3 images were analyzed. Image analysis was carried out with ImageJ.38
Protein and lipid corona formation and characterization
Protein coronas were formed on magNPs, TiO2, SiO2, Ag NPs, and MWCNTs, as previously described.26,27,39,40 In brief, nanomaterials (1 mg mL−1) were incubated with FBS (10437028, Thermo Fisher Scientific, Waltham, MA) at a protein concentration of 40 mg mL−1, L-α-phosphatidylcholine (LAP; 1.8 mg mL−1 in PBS, 44924, Sigma-Aldrich, St. Louis, MO), or FBS supplemented with LAP (1.8 mg mL−1), as described in the text. Experiments also examined coronas formed from bovine serum albumin (BSA; 5 mg ml−1, A2153, Sigma-Aldrich) and transferrin (5 mg mL−1, T8158, Sigma-Aldrich), as well as oleic acid (1.8 mg mL−1, O1257, Sigma-Aldrich), or FBS (40 mg mL−1) supplemented with oleic acid (1.8 mg mL−1). Biomolecules and nanomaterials were combined and then incubated for 30 minutes on a microplate shaker (1000 rpm, 88882006, Digital Microplate Shaker, Thermo Fisher Scientific).To isolate biomolecule–nanomaterial complexes, the mixtures using environmental nanomaterials (TiO2, SiO2, Ag NPs, and MWCNTs) were centrifuged (15 min, 18![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 rcf) and resuspended in PBS three times to remove the majority of unbound and loosely bound biomolecules. When LAP is present, LAP and proteins are pelleted along with the nanomaterials resulting in a centrifugation artifact in the protein assays due to both protein in the pellet and a small false positive from LAP in the protein assay (Fig. S1 and Table S1). The use of oleic acid did not result in this artifact. The treatment of this centrifugation artifact and the protein signal from LAP are described throughout the text.
000 rcf) and resuspended in PBS three times to remove the majority of unbound and loosely bound biomolecules. When LAP is present, LAP and proteins are pelleted along with the nanomaterials resulting in a centrifugation artifact in the protein assays due to both protein in the pellet and a small false positive from LAP in the protein assay (Fig. S1 and Table S1). The use of oleic acid did not result in this artifact. The treatment of this centrifugation artifact and the protein signal from LAP are described throughout the text.
To characterize nanomaterials with only a hard corona and no centrifugation artifact, magNPs were used with magnetic separation of the NPs from the unbound proteins and lipids. Coronas were formed by incubating magNPs (1 mg mL−1) with FBS, LAP, and FBS supplemented with LAP using the same concentrations and conditions described above. The biomolecule–magNP complexes were separated from unbound protein and lipid using magnetic pull-down instead of centrifugation, as described previously.39,40 Centrifuge tubes (89000-028, VWR, Radnor, PA) containing the mixture of biomolecules and magNPs were placed along a magnet (grade 12, 10 mm) for 5 min at room temperature to pull down the biomolecule–magNP complexes. Pull-down and resuspension in PBS was repeated three times (Fig. S2 and Table S2).
Protein and lipid quantification assays were carried out on nanomaterials resuspended in PBS (1 mg mL−1). Protein concentration was measured with a pierce 660 nm protein assay (2260, Thermo Fisher Scientific). For protein assays, protein was eluted from the nanomaterial surface using a 5% solution of sodium dodecyl sulfate (SDS; L3771, Sigma-Aldrich) at 95 °C for 5 min following by centrifugation (5 min, 18![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 rcf) to separate the protein-containing supernatant from the nanomaterial. The limit of detection of the protein assay is 5 μg mL−1 or 5 μg protein per mg nanomaterial. Lipid quantification, measuring unsaturated fatty acids, was performed using a lipid quantification kit (STA-613, Cell Biolabs, San Diego, CA). Lipid corona concentration was measured directly on the magNP surface. The limit of detection of the lipid assay is 50 μg mL−1 or 50 μg lipid per mg nanomaterial.
000 rcf) to separate the protein-containing supernatant from the nanomaterial. The limit of detection of the protein assay is 5 μg mL−1 or 5 μg protein per mg nanomaterial. Lipid quantification, measuring unsaturated fatty acids, was performed using a lipid quantification kit (STA-613, Cell Biolabs, San Diego, CA). Lipid corona concentration was measured directly on the magNP surface. The limit of detection of the lipid assay is 50 μg mL−1 or 50 μg lipid per mg nanomaterial.
Lipidomic analysis of bronchoalveolar lavage fluid (BALF)
BALF samples (n = 6 distinct male volunteers) were obtained from bronchoalveolar lavage (BAL) performed during bronchoscopy in human volunteers.41,42 Prior to BAL, individuals underwent a controlled filtered air exposure with intermittent exercise. A bronchoscopy with BAL in the right middle lobe of the lung was completed ∼21 h following. The lavage return was then processed to remove cells by centrifuge and then stored at −80 °C until use in these studies. BALF samples are maintained at Duke University in the Tighe lab as an active IRB, which allows excess samples to be used for research purposes.Lipidomic analysis of these samples was carried out in the Duke University Proteomics and Metabolomics Core Facility. BALF samples were run on a MxP Quant 500 kit (Biocrates, Innsbruck, Austria) in accordance with the Biocrates protocol. Manufacturer supplied standards and 30 μL of BALF samples were added to the appropriate wells. The plate was then dried under a gentle stream of nitrogen for 30 minutes. The samples were eluted with 5 mM ammonium acetate in methanol. Samples were diluted with a running solvent (a proprietary mixture provided by Biocrates) for flow injection analysis (20![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1). The flow injection analysis-tandem mass spectrometry (FIA-MS/MS) of lipid samples (acylcarnitines, monosaccharides (hexose), diglycerides, triglycerides, lysophosphatidylcholines, phosphatidylcholines, sphingomyelins, ceramides, and cholesteryl esters) was analyzed in two separate FIA-MS/MS methods with total analysis time of approximately 3.8 minutes per injection using (ESI+) on a QTrap 6500+ (Sciex, Marlborough, MA) mass spectrometer. The injection of each sample was performed with the Waters Acquity I-class UPLC (no separation column) at room temperature. The injection volume was 20 μL. The isocratic elution is with a supplier-provided FIA solution. The constant flow rate was 30 μL min−1. Compound-specific precursors to product ion transitions (multiple reaction monitoring (MRM)) for each analyte and the proprietary internal standard were collected. The acquired FIA-MS/MS data were imported into WebIDQ (Biocrates) for peak integration, signal evaluation, calibration curve buildup, and result validation. The top 10 most abundant lipids of the 523 lipids measured were used for further analysis.
1). The flow injection analysis-tandem mass spectrometry (FIA-MS/MS) of lipid samples (acylcarnitines, monosaccharides (hexose), diglycerides, triglycerides, lysophosphatidylcholines, phosphatidylcholines, sphingomyelins, ceramides, and cholesteryl esters) was analyzed in two separate FIA-MS/MS methods with total analysis time of approximately 3.8 minutes per injection using (ESI+) on a QTrap 6500+ (Sciex, Marlborough, MA) mass spectrometer. The injection of each sample was performed with the Waters Acquity I-class UPLC (no separation column) at room temperature. The injection volume was 20 μL. The isocratic elution is with a supplier-provided FIA solution. The constant flow rate was 30 μL min−1. Compound-specific precursors to product ion transitions (multiple reaction monitoring (MRM)) for each analyte and the proprietary internal standard were collected. The acquired FIA-MS/MS data were imported into WebIDQ (Biocrates) for peak integration, signal evaluation, calibration curve buildup, and result validation. The top 10 most abundant lipids of the 523 lipids measured were used for further analysis.
Cell culture
RAW 264.7 mouse macrophages (TIB-71, ATCC, Manassas, VA) were cultured in Dulbecco's modified Eagle's medium (DMEM, 12100046, Thermo Fisher Scientific) supplemented with 10% FBS (F4135, Sigma-Aldrich) at 37 °C and 5% CO2. Cells were passaged by scraping (08100240, Thermo Fisher Scientific) every 2–3 days.Quantification of TNF-α
RAW 264.7 cells were seeded onto sterile 12-well plates (10062-894, VWR) at a density of 700![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 cells per well. The cells were cultured for two days and then incubated with bare nanomaterials (TiO2 NP = 1 mg mL−1; SiO2 NP = 0.1 mg mL−1; Ag NP = 0.25 mg mL−1; MWCNT = 0.05 mg mL−1) with a LAP corona, FBS corona, or FBS + LAP corona. Nanomaterial concentrations were based on prior literature with the goal of eliciting a measurable TNF-α response. Opti-MEM, rather than complete cell culture medium, was used during the 24 hour incubation with nanomaterials to prevent in situ protein corona formation. Experiments were carried out in triplicate. After a 24 hour incubation, the cell media was harvested and centrifuged (15 min, 3000 rcf) to remove any cell debris. The supernatant was collected and stored at −20 °C. TNF-α release was measured using ELISAs according to the manufacturer's instructions (DY410, Mouse TNF-α DuoSet ELISA, R&D Systems, Minneapolis, MN).
000 cells per well. The cells were cultured for two days and then incubated with bare nanomaterials (TiO2 NP = 1 mg mL−1; SiO2 NP = 0.1 mg mL−1; Ag NP = 0.25 mg mL−1; MWCNT = 0.05 mg mL−1) with a LAP corona, FBS corona, or FBS + LAP corona. Nanomaterial concentrations were based on prior literature with the goal of eliciting a measurable TNF-α response. Opti-MEM, rather than complete cell culture medium, was used during the 24 hour incubation with nanomaterials to prevent in situ protein corona formation. Experiments were carried out in triplicate. After a 24 hour incubation, the cell media was harvested and centrifuged (15 min, 3000 rcf) to remove any cell debris. The supernatant was collected and stored at −20 °C. TNF-α release was measured using ELISAs according to the manufacturer's instructions (DY410, Mouse TNF-α DuoSet ELISA, R&D Systems, Minneapolis, MN).
Results and discussion
Four different environmental nanomaterials were used in experiments: TiO2, SiO2, Ag NPs, and MWCNTs (Table 1 and Fig. 1). In addition, magnetic NPs (magNPs) were used as a model system. Hydrodynamic diameter (dh), polydispersity index (PDI), and zeta potential (ZP) were characterized for the roughly spherical TiO2, SiO2, Ag, and magNPs. Primary particle diameter was characterized by TEM (dTEM) for all nanomaterials. The environmental NPs all form aggregates, as has been observed previously, resulting in larger hydrodynamic diameters than primary particle diameters.36,39,43–49 The MWCNT diameter was measured along the short axis. The MWCNT diameter of 12 ± 1.9 nm is in good agreement with previous TEM measurements of the same MWCNTs (11.9 ± 6 nm and 12 nm).27,50| Nanomaterial | Application (supplier) | d h (nm) | PDI | ZP (mV) | d TEM (nm) |
|---|---|---|---|---|---|
| TiO2 | Plastics, paint (Chemours) | 360 ± 14 | 0.3 ± 0.01 | −29 ± 2.9 | 230 ± 84 |
| SiO2 | Construction, biomedical (Millipore Sigma) | 629 ± 82 | 0.5 ± 0.05 | −40 ± 4.5 | 28 ± 7 |
| Ag | Biomedical, antimicrobial (Millipore Sigma) | 764 ± 802 | 0.8 ± 0.2 | −45 ± 3.6 | 126 ± 60 |
| MWCNT | Electronics, construction (Nanocyl) | — | — | — | 12 ± 1.9 |
| MagNP | Model system (Ocean NanoTech) | 205 ± 2 | 0.2 ± 0.04 | −44 ± 2.9 | 129 ± 43 |
 | ||
| Fig. 1 Representative TEM image of TiO2 NPs, SiO2 NPs, Ag NPs, and MWCNTs. The scale bar is 200 nm for all images. | ||
Exogenous LAP adsorbs on magNPs and leads to an increased protein corona
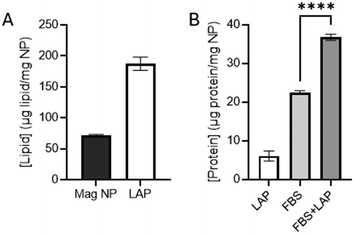
To understand the effect of lipids on the protein corona, we first measured LAP adsorption on magNPs, which we use as a model system. Biomolecule–magNP complexes are prepared using magnetic pull-down instead of centrifugation. This avoids the centrifugation artifact created by lipid and protein pelleting along with the nanomaterials, as described in materials and methods. LAP was selected as a model lipid due to the high abundance of phosphatidylcholines in human BALF (Fig. S3). The concentration of LAP used for experiments (1.8 mg mL−1) was chosen based on adsorption on the surface of magNPs (Fig. S4). Incubation of magNPs (1 mg mL−1) with LAP (1.8 mg mL−1) results in a lipid corona on the magNPs (Fig. 2A). After removing the signal that results from the dye of the lipid assay interacting with the magNPs (70 ± 3 μg lipid per mg NP), we find a LAP concentration of 110 ± 21 μg lipid per mg NP in the corona. This concentration of LAP is 6% of the starting concentration (1.8 mg mL−1) of LAP used to form the corona.By measuring protein concentration, we can examine the effect of LAP on the protein corona. Forming a protein corona from FBS (40 mg mL−1) combined with LAP (1.8 mg mL−1) results in greater protein adsorption than the use of FBS alone (Fig. 2B). Note that a LAP corona alone generates a small positive signal from the protein assay (6 ± 2 μg protein per mg NP). Subtracting this LAP signal from the combined FBS + LAP corona results in a protein signal of 31 ± 3 μg protein per mg NP compared to a FBS corona of 22 ± 1 μg protein per mg NP (p < 0.01). Fig. 2B shows the raw data without the subtraction of the LAP protein signal from the FBS + LAP data. Only a small fraction of FBS forms a corona on the magNPs (0.055%). The addition of LAP increases this to 0.078% of FBS adsorbed on the magNPs.
LAP leads to increased protein in the corona of TiO2, SiO2, Ag NPs and MWCNTs
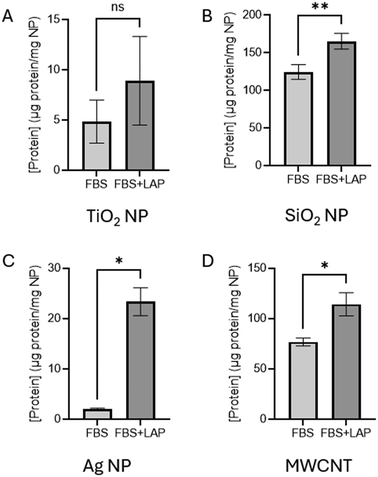
After using magNPs as a model system (Fig. 2), we extended our experiments to TiO2, SiO2, Ag NPs, and MWCNTs (Fig. 3). A similar increase in the concentration of protein adsorbed on the nanomaterials in the presence of LAP was observed, although the percent of that increase varied by nanomaterial (Table 2). Data shown in Fig. 3 includes two modifications. First, as described above for magNPs (Fig. 2B), a corona formed from LAP alone generates a small positive signal from the protein assay (Table S1). That value, which is specific to each type of nanomaterial, is subtracted from the FBS + LAP data shown in Fig. 3. In addition, as described in materials and methods, the artifact resulting from centrifugation with LAP present (Fig. S1; 10 ± 5 μg protein per mg nanomaterial) is subtracted from the FBS + LAP signal. The lipid assay was not compatible with these nanomaterials and lipid corona measurements are not included.| Nanomaterial | FBS adsorbed (%) | Increase with LAP |
|---|---|---|
| TiO2 | 0.012 ± 0.002 | 84% (ns) |
| SiO2 | 0.31 ± 0.024 | 33% |
| Ag | 0.005 ± 0.001 | 1084% |
| MWCNT | 0.19 ± 0.19 | 49% |
This increase in protein corona with the addition of lipid is not universal across lipid types. The addition of oleic acid to FBS did not result in an increased protein corona (Fig. S5).
Individual serum proteins show a nanomaterial-dependent response to coronas formed with LAP present
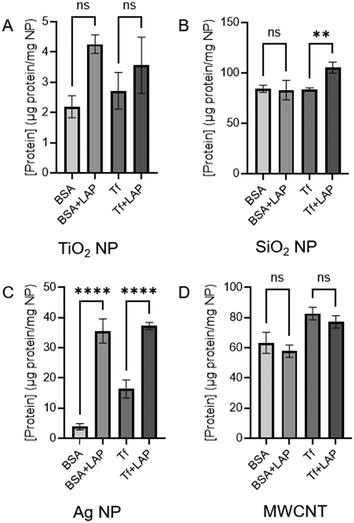
To determine if the increased protein adsorption in the presence of LAP (Fig. 3) was specific to the combination of proteins present in FBS or would also occur for individual serum proteins, we examined two specific serum proteins: bovine serum albumin (BSA) and transferrin. BSA was chosen as the most abundant protein in FBS.39,51–53 Transferrin was selected as a representative blood serum protein that does not directly interact with lipids. We measured the concentration of protein in the corona formed on nanomaterials after incubation with BSA (5 mg mL−1) and transferrin (5 mg mL−1). We find a nanomaterial-dependent response to the adsorption of BSA or transferrin in the presence of LAP (Fig. 4 and Table 3). For example, Ag NPs show a significant increase in both the BSA and transferrin coronas formed in the presence of LAP (Fig. 4C), similar to that observed for the mixture of proteins in FBS (Fig. 3C). MWCNTs show no significant change in the BSA or transferrin coronas with LAP present during corona formation (Fig. 4D). In comparison, the FBS corona on MWCNTs was increased (Fig. 3D), suggesting that the increased FBS adsorption in the presence of LAP is not due to BSA or transferrin, but rather a different serum protein.| Nanomaterial | BSA adsorbed (%) | Increase with LAP | Tf adsorbed (%) | Increase with LAP |
|---|---|---|---|---|
| TiO2 | 0.044 ± 0.013 | 94% (ns) | 0.021 ± 0.054 | 31% (ns) |
| SiO2 | 1.7 ± 0.07 | −1% (ns) | 1.7 ± 0.035 | 26% |
| Ag | 0.078 ± 0.02 | 817% | 0.33 ± 0.06 | 128% |
| MWCNT | 1.3 ± 0.14 | −9% (ns) | 1.7 ± 0.084 | −7% (ns) |
Coronas formed from LAP, FBS, or FBS + LAP result in increased nanomaterial diameter
The formation of a lipid (LAP), protein (FBS), or combined protein–lipid (FBS + LAP) corona on the surface of TiO2, SiO2, and Ag NPs leads to increased hydrodynamic diameter (Table 4). Since MWCNTs are not spherical, they were not included in these comparisons. It should be noted that it is difficult to draw broader conclusions from this DLS data. The nanomaterials, especially with coronas, are highly heterogeneous and aggregated. In addition to the biomolecule–nanomaterial complexes, the LAP and LAP + FBS samples can contain lipid and protein that has been pelleted along with the biomolecule–nanomaterial complex (Fig. S1), which will contribute to the PDI. The one interesting feature to note is the relatively stable PDI for SiO2 NPs incubated with LAP. These samples show a slight decrease in PDI (Δ for LAP = −0.06 ± 0.05 and for FBS + LAP −0.07 ± 0.09). This differentiates the LAP and FBS + LAP coronas from SiO2 NPs with only a FBS corona (Δ = 0.3 ± 0.07) and from the other nanomaterials, which show increased PDI. This suggests that LAP may have a stabilizing effect on the SiO2 NPs, which will be investigated in future studies.| Nanomaterial | Corona | d h (nm) | Δdh (nm) | PDI | ΔPDI | ZP (mV) | ΔZP (mV) |
|---|---|---|---|---|---|---|---|
| TiO2 | LAP | 840 ± 159 | 480 ± 173 | 0.7 ± 0.05 | 0.4 ± 0.05 | −48 ± 1.1 | −19 ± 3 |
| FBS | 1352 ± 50 | 992 ± 52 | 0.9 ± 0.02 | 0.4 ± 0.02 | −32 ± 0.25 | −3 ± 3 | |
| FBS + LAP | 571 ± 15 | 211 ± 21 | 0.5 ± 0.02 | 0.2 ± 0.02 | −38 ± 1.2 | −9 ± 3 | |
| SiO2 | LAP | 1023 ± 51 | 394 ± 97 | 0.4 ± 0.02 | −0.06 ± 0.05 | −32 ± 0.7 | 8 ± 5 |
| FBS | 1162 ± 55 | 533 ± 99 | 0.8 ± 0.05 | 0.3 ± 0.07 | −30 ± 1.6 | 10 ± 5 | |
| FBS + LAP | 979 ± 30 | 350 ± 87 | 0.4 ± 0.07 | −0.07 ± 0.09 | −21 ± 0.1 | 19 ± 5 | |
| Ag | LAP | 1748 ± 300 | 984 ± 856 | 0.9 ± 0.05 | 0.1 ± 0.2 | −41 ± 2.1 | 4 ± 4 |
| FBS | 3526 ± 696 | 2762 ± 1062 | 1 | 0.2 ± 0.2 | −13 ± 0.6 | 32 ± 4 | |
| FBS + LAP | 2834 ± 153 | 2070 ± 816 | 1 | 0.2 ± 0.2 | −31 ± 1 | 14 ± 4 |
LAP in the corona alters TNF-α release from macrophages
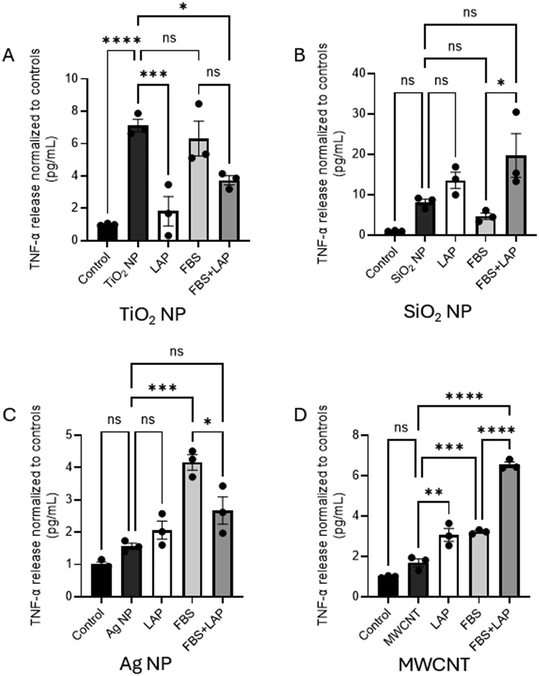
To characterize the cellular response to nanomaterials as a function of LAP in the corona, we used ELISAs to measure the generation of TNF-α, a pro-inflammatory cytokine, in macrophages following a 24 hour incubation with nanomaterials (Fig. 5). TNF-α was chosen as a known indicator of pro-inflammatory cellular response to nanomaterials. Our prior work has shown an increased TNF-α response in macrophages incubated with TiO2 NPs.26,36 We find that LAP present in the corona leads to a nanomaterial-dependent change in the level of TNF-α release. For TiO2 NPs (Fig. 5A), the LAP only (1.8 ± 1.6 pg mL−1) and FBS + LAP (3.7 ± 0.50 pg mL−1) coronas both decrease the TNF-α response compared to bare TiO2 NPs (7.1 ± 0.66 pg mL−1) (Fig. 5A). The increased level of TNF-α release in response to bare TiO2 NPs is in agreement with our previous work showing that bare TiO2 NPs, in the absence of a protein corona, oxidize the plasma membrane of the cell leading to cytotoxicity.54 In comparison, MWCNTs with a LAP (3.07 ± 0.55 pg mL−1) or FBS + LAP (6.55 ± 0.24 pg mL−1) corona lead to an increase in the TNF-α response compared to bare MWCNTs (1.69 ± 0.34 pg mL−1) (Fig. 5D). Cells treated with SiO2 and Ag NPs show little response to the nanomaterials (Fig. 5B and C). SiO2 NPs with a FBS + LAP corona have an increased TNF-α response compared to FBS (Fig. 5B). Ag NPs with a FBS + LAP corona have a decreased TNF-α response compared to FBS (Fig. 5C).The cellular response to nanomaterials is complex and there is no reason to expect similar trends across TiO2, SiO2, Ag NPs and MWCNTs. The nanomaterial concentrations used for the 24 hour incubation are not matched (TiO2 NP = 1 mg mL−1; SiO2 NP = 0.1 mg mL−1; Ag NP = 0.25 mg mL−1; MWCNT = 0.05 mg mL−1) and were chosen to elicit a TNF-α response. It is also important to note that FBS contains endogenous lipids (1.08 ± 0.09 mg mL−1, Table S3) and an increase in the FBS corona may also include an increase in the concentration and types of lipids in the corona beyond LAP. Our own previous work has shown that TiO2 NP aggregation is an important driver in cytokine response.36 Ag NPs may undergo dissolution generating ions and also altering particle features during the 24 hour incubation period (Fig. S6). Future experiments will need to determine how the LAP, FBS, FBS + LAP coronas alter the interaction of these nanomaterials with cells. For example, previous work with polystyrene NPs (∼100 nm) and macrophages (RAW 264.7) has shown, using flow cytometry, decreased cellular association of polystyrene NPs with a lipoprotein corona, compared to bare polystyrene NPs. A corona formed only from plasma, similar to our FBS corona, had the lowest level of cellular association.55 In comparison, confocal fluorescence microscopy experiments with magnetite nanoparticles, with either hydrophilic (starch) or hydrophobic (phosphatidylcholine) surface modifications, and alveolar macrophages (MH-S) showed that surfactant lipids, rather than proteins, dominate the cellular response.56 Flow cytometry experiments with TiO2 NPs and alveolar macrophages found high levels of TiO2 NP uptake with only modest increases in the presence of surfactant.57 It is possible that the altered TNF-α response that we observe is due to a combination of lipid and protein–nanomaterial interactions as well as an altered interaction of the nanomaterial with macrophages.
Conclusions
Our experiments examine the interaction of four industrially- and environmentally-relevant nanomaterials; TiO2, SiO2, and Ag NPs and MWCNTs (Table 1 and Fig. 1) with lipids, proteins, and cells. We find that the presence of a representative lung lining fluid lipid (LAP) during corona formation leads to an increased concentration of FBS proteins adsorbed on the surface of these nanomaterials (Fig. 2 and 3, Table 2), along with increased hydrodynamic diameter (Table 4). The adsorption of two individual serum proteins, BSA and transferrin, is nanomaterial-dependent (Fig. 4 and Table 3). The presence of LAP in the corona alters the cellular response measured as release of TNF-α, a pro-inflammatory cytokine (Fig. 5). The specific response to LAP, FBS, and FBS + LAP coronas is dependent on the specific nanomaterial. In the case of TiO2 NPs, LAP provides a protective effect (Fig. 5A).Researchers have proposed lipid corona “fingerprinting” in parallel with the existing detailed studies of the protein corona.33 However, previous work has suggested that the lipid profile is independent of nanomaterial surface properties.34 In addition, coarse-grained molecular dynamics simulations of Ag (5 nm, 15 nm) and polystyrene (5 nm, 10 nm) NPs showed that hydrophobicity was a key factor in the adsorption of pulmonary surfactant proteins on the NP surface, but the lipid composition of the corona was independent of NP properties.58 Our work did not identify specific lipids present on the nanomaterial surface, but does demonstrate that lipids are an important component of the protein corona and cellular response. Instead, our experiments used LAP as a representative lipid based on the abundance of phosphatidylcholines in lung lining fluid (Fig. S3). Previous work has shown that phosphatidylcholine-iron oxide micelles (75 nm) improved circulation times in rats compared to PEG-functionalized NPs (25 nm).59 Proteomic analysis suggested that the specific proteins adsorbed on the phosphatidylcholine-functionalized micelles presented an anti-fouling surface that increased circulation. As our work did not probe the specific composition of the protein corona, only the concentration (Fig. 2–4), this provides an interesting next step for comparison.
Beyond these fundamental studies, the interaction of lipids with nanomaterials and the resulting changes in the protein corona could have important clinical implications. Previous work has shown that the protein corona is altered by high cholesterol levels in mouse serum and that this change in the protein corona changes cellular uptake and biodistribution to individual organs.60 Our research highlights the importance of controlled molecular studies to understand complex cellular responses to nanomaterials with potential clinical implications.
Author contributions
M. M. H., R. M. T., and C. K. P. conceived the project. M. M. H., C. C., A. K., and E. M. S. conducted experiments. M. M. H., C. C., and A. K. analyzed results. M. M. H. and C. K. P. wrote the manuscript. All authors reviewed the manuscript.Conflicts of interest
The authors declare that they have no conflicts of interest.Data availability
All study data are included in the article and/or the supplementary information (SI).Supplementary information is available. See DOI: https://doi.org/10.1039/d5en00971e.
Acknowledgements
This study received funding from the NIH-NIEHS (5R01ES032443) and Duke University MEDx. We thank the Duke University School of Medicine for the use of the Proteomics and Metabolomics Core Facility, which provided lipidomic services. We thank Michela Albright for assistance with experiments.References
- A. Mohajerani, L. Burnett, J. Smith, H. Kurmus, J. Milas, A. Arulrajah, S. Horpibulsuk and A. A. Kadir, Nanoparticles in construction materials and other applications, and implications of nanoparticle use, Materials, 2019, 12, 3052–3077 CrossRef CAS PubMed.
- M. J. Hanus and A. T. Harris, Nanotechnology innovations for the construction industry, Prog. Mater. Sci., 2013, 58, 1056–1102 CrossRef CAS.
- J. Lee, S. Mahendra and P. J. J. Alvarez, Nanomaterials in the construction industry: A review of their applications and environmental health and safety considerations, ACS Nano, 2010, 4, 3580–3590 CrossRef CAS PubMed.
- S. F. Hansen, O. F. H. Hansen and M. B. Nielsen, Advances and challenges towards consumerization of nanomaterials, Nat. Nanotechnol., 2020, 15, 964–965 CrossRef CAS PubMed.
- R. Kessler, Engineered nanoparticles in consumer products: Understanding a new ingredient, Environ. Health Perspect., 2011, 119, 120–125 Search PubMed.
- A. Weir, P. Westerhoff, L. Fabricius, K. Hristovski and N. Von Goetz, Titanium dioxide nanoparticles in food and personal care products, Environ. Sci. Technol., 2012, 46, 2242–2250 CrossRef CAS PubMed.
- V. Kodali, E. Gyung Lee, R. Gill, A. Afshari, W. McKinney, G. Casuccio, K. Bunker, T. Lersch, K. Rickabaugh and A. Erdely, 111 toxicity assessment of a carbon nanotube embedded concrete, Ann. Work Exposures Health, 2023, 67, i64–i65 CrossRef.
- K. E. Wheeler, A. J. Chetwynd, K. M. Fahy, B. S. Hong, J. A. Tochihuitl, L. A. Foster and I. Lynch, Environmental dimensions of the protein corona, Nat. Nanotechnol., 2021, 16, 617–629 CrossRef CAS PubMed.
- Z. Zahra, Z. Habib, S. Chung and M. A. Badshah, Exposure route of TiO2 NPs from industrial applications to wastewater treatment and their impacts on the agro-environment, Nanomater., 2020, 10, 1469 CrossRef CAS PubMed.
- S. Runa, M. Hussey and C. K. Payne, Nanoparticle–cell interactions: Relevance for public health, J. Phys. Chem. B, 2018, 122, 1009–1016 CrossRef CAS PubMed.
- M. G. Soliman, A. Martinez-Serra, G. Antonello, M. Dobricic, T. Wilkins, T. Serchi, I. Fenoglio and M. P. Monopoli, Understanding the role of biomolecular coronas in human exposure to nanomaterials, Environ. Sci.: Nano, 2024, 11, 4421–4448 RSC.
- M. M. Nabi, J. Wang, M. Erfani, E. Goharian and M. Baalousha, Urban runoff drives titanium dioxide engineered particle concentrations in urban watersheds: field measurements, Environ. Sci.: Nano, 2023, 10, 718–731 RSC.
- J. Zhang, W. Guo, Q. Li, Z. Wang and S. Liu, The effects and the potential mechanism of environmental transformation of metal nanoparticles on their toxicity in organisms, Environ. Sci.: Nano, 2018, 5, 2482–2499 RSC.
- S. Lin, M. Mortimer, R. Chen, A. Kakinen, J. E. Riviere, T. P. Davis, F. Ding and P. C. Ke, NanoEHS beyond toxicity – focusing on biocorona, Environ. Sci.: Nano, 2017, 4, 1433–1454 RSC.
- U.S. Department of Labor, OSHA fact sheet: Working safely with nanomaterials, 2023.
- D. J. You and J. C. Bonner, Susceptibility factors in chronic lung inflammatory responses to engineered nanomaterials, Int. J. Mol. Sci., 2020, 21, 7310–7339 CrossRef CAS PubMed.
- H. Shi, R. Magaye, V. Castranova and J. Zhao, Titanium dioxide nanoparticles: A review of current toxicological data, Part. Fibre Toxicol., 2013, 5, 1–33 Search PubMed.
- M. Falahati, A. Hasan, H. A. Zeinabad, V. Serpooshan, J. H. Von Der Thüsen and T. L. M. Ten Hagen, Engineering of pulmonary surfactant corona on inhaled nanoparticles to operate in the lung system, Nano Today, 2023, 52, 101998–102015 CrossRef CAS.
- S. Hussain, J. A. J. Vanoirbeek and P. H. M. Hoet, Interactions of nanomaterials with the immune system, Wiley Interdiscip. Rev.: Nanomed. Nanobiotechnol., 2012, 4, 169–183 CAS.
- D. Pelclova, V. Zdimal, Z. Fenclova, S. Vlckova, F. Turci, I. Corazzari, P. Kacer, J. Schwarz, N. Zikova, O. Makes, K. Syslova, M. Komarc, J. Belacek, T. Navratil, M. Machajova and S. Zakharov, Markers of oxidative damage of nucleic acids and proteins among workers exposed to TiO2 (nano) particles, Occup. Environ. Med., 2016, 73, 110–118 CrossRef CAS PubMed.
- B. Sha, W. Gao, X. Cui, L. Wang and F. Xu, The potential health challenges of TiO2 nanomaterials, J. Appl. Toxicol., 2015, 35, 1086–1101 CrossRef CAS PubMed.
- M. Geiser and W. G. Kreyling, Deposition and biokinetics of inhaled nanoparticles, Part. Fibre Toxicol., 2010, 7, 2–19 CrossRef PubMed.
- D. Nierenberg, A. R. Khaled and O. Flores, Formation of a protein corona influences the biological identity of nanomaterials, Rep. Pract. Oncol. Radiother., 2018, 23, 300–308 CrossRef PubMed.
- A. J. Chetwynd and I. Lynch, The rise of the nanomaterial metabolite corona, and emergence of the complete corona, Environ. Sci.: Nano, 2020, 7, 1041–1060 RSC.
- N. V. Konduru, R. M. Molina, A. Swami, F. Damiani, G. Pyrgiotakis, P. Lin, P. Andreozzi, T. C. Donaghey, P. Demokritou, S. Krol, W. Kreyling and J. D. Brain, Protein corona: implications for nanoparticle interactions with pulmonary cells, Part. Fibre Toxicol., 2017, 14, 42–54 CrossRef PubMed.
- K. Poulsen, M. Albright, N. J. Niemuth, R. M. Tighe and C. K. Payne, Interaction of TiO2 nanoparticles with lung fluid proteins and the resulting macrophage inflammatory response, Environ. Sci.: Nano, 2023, 10, 2427–2437 RSC.
- J. Dominguez, S. K. Holmes, R. D. Bartone, L. J. Tisch, R. M. Tighe, J. C. Bonner and C. K. Payne, House dust mite extract forms a der p 2 corona on multi-walled carbon nanotubes: implications for allergic airway disease, Environ. Sci.: Nano, 2024, 11, 324–335 RSC.
- A. Holian, R. F. Hamilton, Z. Wu, S. Deb, K. L. Trout, Z. Wang, R. Bhargava and S. Mitra, Lung deposition patterns of MWCNT vary with degree of carboxylation, Nanotoxicology, 2019, 13, 143–159 CrossRef CAS PubMed.
- R. Leibe, I.-L. Hsiao, S. Fritsch-Decker, U. Kielmeier, A. M. Wagbo, B. Voss, A. Schmidt, S. D. Hessman, A. Duschl, G. J. Oostingh, S. Diabaté and C. Weiss, The protein corona suppresses the cytotoxic and pro-inflammatory response in lung epithelial cells and macrophages upon exposure to nanosilica, Arch. Toxicol., 2019, 93, 871–885 CrossRef CAS PubMed.
- C. W. Agudelo, G. Samaha and I. Garcia-Arcos, Alveolar lipids in pulmonary disease A review, Lipids Health Dis., 2020, 19, 122–143 CrossRef CAS PubMed.
- R. Veldhuizen, K. Nag, S. Orgeig and F. Possmayer, The role of lipids in pulmonary surfactant, Biochim. Biophys. Acta, Mol. Basis Dis., 1998, 1408, 90–108 CrossRef CAS PubMed.
- T. Lima, K. Bernfur, M. Vilanova and T. Cedervall, Understanding the lipid and protein corona formation on different sized polymeric nanoparticles, Sci. Rep., 2020, 10, 1129–1138 CrossRef CAS PubMed.
- X. Zhang, A. K. Pandiakumar, R. J. Hamers and C. J. Murphy, Quantification of lipid corona formation on colloidal nanoparticles from lipid vesicles, Anal. Chem., 2018, 90, 14387–14394 CrossRef CAS PubMed.
- S. S. Raesch, S. Tenzer, W. Storck, A. Rurainski, D. Selzer, C. A. Ruge, J. Perez-Gil, U. F. Schaefer and C.-M. Lehr, Proteomic and lipidomic analysis of nanoparticle corona upon contact with lung surfactant reveals differences in protein, but not lipid composition, ACS Nano, 2015, 9, 11872–11885 CrossRef CAS PubMed.
- X. Bai, S. M. Lam, P. Nie, M. Xu, S. Liu, G. Shui and G. Hu, Lipidomic analysis probes lipid coronas on hydrophilic nanoparticles from natural lung surfactant, Environ. Sci.: Nano, 2022, 9, 4150–4161 RSC.
- M. M. Heckman, M. C. Albright, K. M. Poulsen, R. M. Tighe and C. K. Payne, Cellular and In Vivo response to industrial, food grade, and photocatalytic TiO2 nanoparticles, J. Phys. Chem. B, 2024, 128, 8878–8885 CrossRef CAS PubMed.
- J. Clogston and A. Vermilya, NCL method PCC-2: Measuring zeta potential of nanoparticles, Nanotechnol. Charact. Lab., 2020, vol. 1.2, pp. 1–13 Search PubMed.
- C. A. Schneider, W. S. Rasband and K. W. Eliceiri, NIH Image to ImageJ: 25 years of image analysis, Nat. Methods, 2012, 9, 671–675 CrossRef CAS PubMed.
- K. Poulsen and C. K. Payne, Concentration and composition of the protein corona as a function of incubation time and serum concentration: An automated approach to the protein corona, Anal. Bioanal. Chem., 2022, 414, 7265–7275 CrossRef CAS PubMed.
- K. Poulsen, T. Pho, J. A. Champion and C. K. Payne, Automation and low-cost proteomics for characterization of the protein corona: Experimental methods for big data, Anal. Bioanal. Chem., 2020, 412, 6543–6551 CrossRef CAS PubMed.
- M. A. Guttenberg, A. T. Vose, A. Birukova, K. Lewars, R. I. Cumming, M. C. Albright, J. I. Mark, C. J. Salazar, S. Swaminathan, Z. Yu, Y. V. Sokolenko, E. Bunyan, M. J. Yaeger, M. B. Fessler, L. G. Que, K. M. Gowdy, A. V. Misharin and R. M. Tighe, Tissue-resident alveolar macrophages reduce ozone-induced inflammation via MerTK-mediated efferocytosis, Am. J. Respir. Cell Mol. Biol., 2024, 70, 493–506 CrossRef CAS PubMed.
- B. W. Frush, Z. Li, J. V. Stiles, S. F. Cotter, S. L. Shofer, W. M. Foster, J. W. Hollingsworth and R. M. Tighe, Ozone primes alveolar macrophage–derived innate immunity in healthy human subjects, J. Allergy Clin. Immunol., 2016, 138, 1213–1215.e1 CrossRef CAS PubMed.
- X. Liu, Y. Xue, T. Ding and J. Sun, Enhancement of proinflammatory and procoagulant responses to silica particles by monocyte-endothelial cell interactions, Part. Fibre Toxicol., 2012, 9, 36–51 CrossRef CAS PubMed.
- D. Breznan, D. D. Das, J. S. O'Brien, C. MacKinnon-Roy, S. Nimesh, N. Q. Vuong, S. Bernatchez, N. DeSilva, M. Hill, P. Kumarathasan and R. Vincent, Differential cytotoxic and inflammatory potency of amorphous silicon dioxide nanoparticles of similar size in multiple cell lines, Nanotoxicology, 2017, 11, 223–235 CrossRef CAS PubMed.
- L. Rubio, G. Pyrgiotakis, J. Beltran-Huarac, Y. Zhang, J. Gaurav, G. Deloid, A. Spyrogianni, K. A. Sarosiek, D. Bello and P. Demokritou, Safer-by-design flame-sprayed silicon dioxide nanoparticles: the role of silanol content on ROS generation, surface activity and cytotoxicity, Part. Fibre Toxicol., 2019, 16, 40–55 CrossRef PubMed.
- M. Xiang, C. Chen, Y. Chen, Y. Zhang, L. Shi, Y. Chen, J. Li, B. Li, B. Zeng, H. R. Xing, J. Wang and Z. Zou, Unexpected inhibitory role of silica nanoparticles on lung cancer development by promoting M1 polarization of macrophages, Int. J. Nanomed., 2024, 19, 11087–11104 CrossRef CAS PubMed.
- L.-S. Gerber, H. J. Heusinkveld, C. Langendoen, B. Stahlmecke, R. P. Schins and R. H. Westerink, Acute, sub-chronic and chronic exposures to TiO2 and Ag nanoparticles differentially affects neuronal function in vitro, NeuroToxicology, 2022, 93, 311–323 CrossRef CAS PubMed.
- K. Sheehy, A. Casey, A. Murphy and G. Chambers, Antimicrobial properties of nano-silver: A cautionary approach to ionic interference, J. Colloid Interface Sci., 2015, 443, 56–64 CrossRef CAS PubMed.
- J. Liu, Z. Huang, S. Yin, X. Zhou, Y. Jiang and L. Shao, The lysosome-mitochondrion crosstalk engaged in silver nanoparticles-disturbed mitochondrial homeostasis, Sci. Total Environ., 2023, 889, 164078–164090 CrossRef CAS PubMed.
- A. J. Taylor-Just, M. D. Ihrie, K. S. Duke, H. Y. Lee, D. J. You, S. Hussain, V. K. Kodali, C. Ziemann, O. Creutzenberg, A. Vulpoi, F. Turcu, M. Potara, M. Todea, S. Van Den Brule, D. Lison and J. C. Bonner, The pulmonary toxicity of carboxylated or aminated multi-walled carbon nanotubes in mice is determined by the prior purification method, Part. Fibre Toxicol., 2020, 17, 1–18 CrossRef PubMed.
- N. C. Cady, A. D. Strickland and C. A. Batt, Optimized linkage and quenching strategies for quantum dot molecular beacons, Mol. Cell. Probes, 2007, 21, 116–124 CrossRef CAS PubMed.
- R. Pieper, C. L. Gatlin, A. J. Makusky, P. S. Russo, C. R. Schatz, S. S. Miller, Q. Su, A. M. McGrath, M. A. Estock, P. P. Parmar, M. Zhao, S. Huang, J. Zhou, F. Wang, R. Esquer-Blasco, N. L. Anderson, J. Taylor and S. Steiner, The human serum proteome: Display of nearly 3700 chromatographically separated protein spots on two-dimensional electrophoresis gels and identification of 325 distinct proteins, Proteomics, 2003, 3, 1345–1364 CrossRef CAS PubMed.
- N. L. Anderson and N. G. Anderson, The human plasma proteome, Mol. Cell. Proteomics, 2002, 1, 845–867 CrossRef CAS PubMed.
- S. Runa, D. Khanal, M. L. Kemp and C. K. Payne, TiO2 nanoparticles alter the expression of peroxiredoxin antioxidant genes, J. Phys. Chem. C, 2016, 120, 20736–20742 CrossRef CAS.
- J. Müller, D. Prozeller, A. Ghazaryan, M. Kokkinopoulou, V. Mailänder, S. Morsbach and K. Landfester, Beyond the protein corona – lipids matter for biological response of nanocarriers, Acta Biomater., 2018, 71, 420–431 CrossRef PubMed.
- C. A. Ruge, U. F. Schaefer, J. Herrmann, J. Kirch, O. Cañadas, M. Echaide, J. Pérez-Gil, C. Casals, R. Müller and C.-M. Lehr, The interplay of lung surfactant proteins and lipids assimilates the macrophage clearance of nanoparticles, PLoS One, 2012, 7, e40775–e40785 CrossRef CAS PubMed.
- B. Stringer and L. Kobzik, Alveolar macrophage uptake of the environmental particulate titanium dioxide: role of surfactant components, Am. J. Respir. Cell Mol. Biol., 1996, 14, 155–160 CrossRef CAS PubMed.
- Q. Hu, X. Bai, G. Hu and Y. Y. Zuo, Unveiling the molecular structure of pulmonary surfactant corona on nanoparticles, ACS Nano, 2017, 11, 6832–6842 CrossRef CAS PubMed.
- H. Groult, J. Ruiz-Cabello, A. V. Lechuga-Vieco, J. Mateo, M. Benito, I. Bilbao, M. P. Martínez-Alcázar, A. Juan, J. Vázquez Lopez and F. F. Herranz, Phosphatidylcholine-coated iron oxide nanomicelles for In Vivo prolonged circulation time with an antibiofouling protein corona, Chem. – Eur. J., 2014, 20, 16662–16671 CrossRef CAS PubMed.
- H. Tang, Y. Zhang, T. Yang, C. Wang, Y. Zhu, L. Qiu, J. Liu, Y. Song, L. Zhou, J. Zhang, Y. K. Wong, Y. Liu, C. Xu, H. Wang and J. Wang, Cholesterol modulates the physiological response to nanoparticles by changing the composition of protein corona, Nat. Nanotechnol., 2023, 18, 1067–1077 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |