Designing well-defined photopolymerized synthetic matrices for three-dimensional culture and differentiation of induced pluripotent stem cells†
Elisa M.
Ovadia
a,
David W.
Colby
a and
April M.
Kloxin
 *ab
*ab
aDepartment of Chemical and Biomolecular Engineering, University of Delaware, Newark, DE 19716, USA. E-mail: akloxin@udel.edu; Fax: +1 (302) 831-1048; Tel: +1 (302) 831-3009
bDepartment of Material Science and Engineering, University of Delaware, Newark, DE 19716, USA
First published on 10th April 2018
Abstract
Induced pluripotent stem cells (iPSCs) are of interest for the study of disease, where these cells can be derived from patients and have the potential to be differentiated into any cell type; however, three-dimensional (3D) culture and differentiation of iPSCs within well-defined synthetic matrices for these applications remains limited. Here, we aimed to establish synthetic cell-degradable hydrogels that allow precise presentation of specific biochemical cues for 3D culture of iPSCs with relevance for hypothesis testing and lineage-specific differentiation. We synthesized poly(ethylene glycol)-(PEG)-peptide-based hydrogels by photoinitiated step growth polymerization and used them to test the hypothesis that the viability of iPSCs within these matrices could be rescued with appropriate biochemical cues inspired by proteins and integrins important for iPSC culture on Matrigel. Specifically, we selected a range of motifs inspired by iPSC binding to Matrigel, including laminin-derived IKVAV and YIGSR, α5β1-binding PHSRNG10RGDS, αvβ5-binding KKQRFRHRNRKG, and RGDS that is known to bind a variety of integrins for generally promoting cell adhesion. YIGSR and PHSRNG10RGDS resulted in the highest iPSC viability, where binding of β1 integrin was key, and these permissive compositions also allowed iPSC differentiation into neural progenitor cells (NPCs) (decreased oct4 expression and increased pax6 expression) in response to soluble factors. The resulting NPCs formed clusters of different sizes in response to each peptide, suggesting that matrix biochemical cues affect iPSC proliferation and clustering in 3D culture. In summary, we have established photopolymerizable synthetic matrices for the encapsulation, culture, and differentiation of iPSCs for studies of cell–matrix interactions and deployment in disease models.
1. Introduction
Pluripotent stem cells, such as embryonic stem cells (ESCs) and induced pluripotent stem cells (iPSCs), are of continued and growing interest for the creation of model systems to study human disease.1,2 For example, iPSCs can be generated from patient-derived cells and differentiated down a number of lineages, including ectoderm, endoderm, and mesoderm, for use in fundamental biological studies and development of disease models for drug screening.1,3,4 In these applications, iPSCs typically have been cultured on top of either an inactivated mouse embryonic fibroblast feeder layer or a thin layer of Matrigel coated on tissue culture treated plastic (TCP).5 While useful, these culture systems have inherent batch-to-batch variability, where Matrigel is harvested from Engelbreth–Holm–Swarm (EHS) mouse sarcoma, a tumor enriched in extracellular matrix proteins found in basement membrane.6 Further, the planar nature of these culture systems polarizes cells in the z-direction, unlike the multidimensional in vivo environment in which cells are surrounded by their microenvironment and related cues (e.g., cell–cell and cell–matrix interactions). To reduce culture variability, TCP coated with whole proteins (i.e., collagen-IV, laminin, fibronectin, vitronectin) or recombinant protein isoforms of them has been shown to promote stem cell pluripotency, self-renewal, and differentiation for two-dimensional (2D) culture;7–11 however, three-dimensional (3D) culture has been limited. Well-defined materials with tunable properties relevant for 3D culture of cells provide opportunities to probe which factors may play a role in iPSC viability and differentiation while providing a multidimensional environment.Several synthetic matrices have been utilized for the culture and expansion of pluripotent stem cells in three dimensions. In particular, hydrogels, crosslinked water-absorbing polymer networks, have been used to mimic key properties of the extracellular matrix (ECM) found within different in vivo niches for stem cell culture, including stem cell expansion and studies of their self-renewal, pluripotency, and differentiation.12–15 For example, a thermoreversible hydrogel (poly(N-isopropylacrylamide)-co-poly(ethylene glycol), PNIPAAm-PEG) was designed for 3D culture and long-term expansion (∼1027-fold over 280 days) of a variety of multipotent and pluripotent stem cells broadly for biomedical applications, including neural differentiation.16 Additionally, in this seminal study by Lei et al., inclusion of a ROCK inhibitor in the culture medium, which prevents apoptosis of dissociated stem cells, was identified as important for stem cell expansion within these polymer networks.16 Other investigations have evaluated the effect of cell–matrix and cell–cell properties on ESC proliferation and self-renewal within hydrogel-based matrices.17,18 For example, 3D hydrogel microarrays formed with Michael-type thiol–ene reactions have been used for high-throughput screening of the effects of multiple extracellular factors on ESC proliferation and self-renewal: soluble factors (leukemia inhibitory factor), degradability of the hydrogel, and low matrix modulus were found to be most influential. In a complementary study, integrin-binding peptides were presented within a similar system to evaluate their influence on self-renewal and stemness of ESCs.16,19,20
While a number of 3D culture studies have examined how cell–cell and cell–matrix interactions influence pluripotency and self-renewal, another important property of pluripotent cells is their potential to differentiate down a variety of lineages for the generation of progenitor and terminally-differentiated cell subtypes.1,21 For example, iPSCs can be generated from cells derived from patients for studies of both regeneration and disease, from mechanistic investigations to drug screening,22,23 where many are particularly interested in iPSC differentiation into neural subtypes for studying neurodegenerative diseases.24 However, to date, few studies in well-defined synthetic matrices have focused on the encapsulation and subsequent differentiation of iPSCs,16 which could prove useful in such disease modeling applications. Progenitor cells that were derived from iPSCs have been encapsulated and cultured in hydrogels, where conditions have been optimized for progenitor cell growth and differentiation. For example, biomimetic peptides have been incorporated within an acrylated hyaluronic-acid hydrogel, formed by Michael-type addition with cysteine-functionalized peptides, where an optimized peptide cocktail enhanced neuronal differentiation of neural progenitor cells (NPCs) derived from iPSCs in 3D culture.25
Despite these advances, many researchers utilize Matrigel for the culture of pluripotent stem cells, partly owing to the challenge of maintaining iPSC viability in many synthetic culture systems while also permitting complex downstream differentiation processes. This harvested material contains a mixture of both large and small bioactive proteins, including laminin, collagen IV, and entactin with some batch-to-batch variation, and has been shown to bind key integrins on iPSCs, including α5, α6, αv, β1, and β5.26,27 Additionally, ROCK inhibition has been observed to increase the expression of these integrins (αv, α6, β1) during iPSC culture on Matrigel.28 We hypothesized that incorporation of ligands that bind to integrins key for iPSC adhesion to and survival on Matrigel within a well-defined synthetic biomaterial would permit iPSC culture, growth, and differentiation in three dimensions.
To test this, we sought to establish a 3D culture platform utilizing light-mediated chemistries for probing key cues for maintaining iPSC viability and enabling their differentiation. Light initiated ‘click’ chemistries allow for rapid, triggered formation of hydrogels, enabling the encapsulation of single cells, and inclusion of specific biochemical cues often independent of the mechanical properties of the network.29,30 As has been demonstrated by our lab and others, this mechanism of hydrogel formation affords spatio-temporal control to incorporate different cues based on where and when light is applied.31–35 Such materials have been used for the encapsulation and culture of multipotent or iPSC-derived cells. For example, McKinnon and co-workers have used photo-initiated hydrogels for controlled encapsulation of ES-derived motor neurons (ESMNs) and, in a follow up study, demonstrated the effect of controlled photo-degradation for the formation of channels to promote axon extension and connectivity, including with other cell types.36,37 However, to date, there are few demonstrations of the use of light-mediated chemistries with pluripotent cells.
In this work, we aimed to establish an approach for the encapsulation and culture of iPSCs within photoinitiated PEG-peptide-based synthetic matrices and to utilize this system to test hypotheses about key cell–matrix interactions for both maintaining iPSC viability and permitting their differentiation. Specifically, we designed poly(ethylene glycol) (PEG)-based hydrogels formed by light-triggered thiol–ene ‘click’ chemistry with specific receptor-binding peptides for investigating their influence on iPSC viability and differentiation. We encapsulated a single-cell suspension of iPSCs and tested the effect of ROCK inhibition and integrin binding on iPSC viability in a cell-degradable hydrogel initially presenting RGDS, which is known to bind a number of integrins for promoting cell adhesion.38 We then selected a range of motifs inspired by iPSC binding to Matrigel, including laminin-derived IKVAV and YIGSR, α5β1-binding PHSRNG10RGDS, and αvβ5-binding KKQRFRHRNRKG, and assessed their effect on iPSC viability and proliferation in 3D culture. In these studies, both ROCK inhibition and activation of β1 integrin were observed to be key for iPSC viability in 3D culture within these synthetic matrices. Lastly, we examined if these environments were not only permissive to iPSC viability and growth but also their pluripotency and subsequent differentiation in the presence of appropriate soluble factors. As pluripotent stem cells have become increasingly used for a variety of biological applications, this approach and the resulting synthetic matrices provide useful tools for future investigations for testing hypotheses about key cell–matrix interactions in both fundamental and translation studies and providing multidimensional platforms for disease culture models and drug screening.39–41
2. Experimental
2.1. Synthesis and characterization of norbornene-functionalized PEG
Multi-arm PEG was functionalized with norbornene end groups according to established protocols.42 8-arm PEG-norbornene (PEG-8-Nb, Mn ∼ 40 kDa) was synthesized by dissolving PEG-8-NH2·HCl (5 g, Jenkem) in 25 mL of anhydrous N,N-dimethylformamide (DMF; ThermoFisher) stirred at room temperature with a magnetic stir bar in a 250 mL round bottom flask. In another 250 mL round bottom flask, 5-norbornene-2-carboxylic acid (Nb-COOH) (2.2 excess relative to amine end groups on PEG; 17.6 molar equivalent) (Sigma Aldrich; St Louis, MO), HATU (2 excess; 16 molar equivalent) (Sigma-Aldrich; St Louis, MO), and 4-methylmorpholine (4-MMP) (4.5 excess; 36 molar equivalent) (Sigma Aldrich; St Louis, MO) were dissolved in 10 mL of DMF in a 250 mL round bottom flask stirring at room temperature. After all components were dissolved, they were combined into one flask (total volume of ∼35 mL) and allowed to react overnight stirring at room temperature. The next day the solution was precipitated twice in ice cold diethyl ether (500 mL, 14× excess diethyl ether relative to DMF) followed by filtration through a Buchner funnel with filter paper to recover the precipitated polymer product. After two precipitations and filtrations, the solid PEG product was allowed to dry in the fume hood overnight. Finally, the PEG product was purified by dialysis (MWCO 2000 g mol−1, Spectrum Laboratories) against continuously-stirred deionized water for 48 hours. Product purity was confirmed by 1H NMR in DMSO-d6: 400 mH δ 6.20 to 5.86 (m, 2H), δ 3.65 to 3.40 (m, 454H), and norbornene functionality was determined to be 92% (ESI Fig. S1†). PEG-8-Nb was stored at −20 °C after lyophilization.2.2. Synthesis and characterization of peptides
The crosslinking peptide (GCRD![[V with combining low line]](https://www.rsc.org/images/entities/char_0056_0332.gif)
![[P with combining low line]](https://www.rsc.org/images/entities/char_0050_0332.gif)
![[M with combining low line]](https://www.rsc.org/images/entities/char_004d_0332.gif)
![[S with combining low line]](https://www.rsc.org/images/entities/char_0053_0332.gif) ↓
↓![[M with combining low line]](https://www.rsc.org/images/entities/char_004d_0332.gif)
![[R with combining low line]](https://www.rsc.org/images/entities/char_0052_0332.gif)
![[G with combining low line]](https://www.rsc.org/images/entities/char_0047_0332.gif)
![[G with combining low line]](https://www.rsc.org/images/entities/char_0047_0332.gif) DRCG) and pendant peptides (CGKG
DRCG) and pendant peptides (CGKG![[Y with combining low line]](https://www.rsc.org/images/entities/char_0059_0332.gif)
![[I with combining low line]](https://www.rsc.org/images/entities/char_0049_0332.gif)
![[G with combining low line]](https://www.rsc.org/images/entities/char_0047_0332.gif)
![[S with combining low line]](https://www.rsc.org/images/entities/char_0053_0332.gif)
![[R with combining low line]](https://www.rsc.org/images/entities/char_0052_0332.gif) , CG
, CG![[K with combining low line]](https://www.rsc.org/images/entities/char_004b_0332.gif)
![[K with combining low line]](https://www.rsc.org/images/entities/char_004b_0332.gif)
![[Q with combining low line]](https://www.rsc.org/images/entities/char_0051_0332.gif)
![[R with combining low line]](https://www.rsc.org/images/entities/char_0052_0332.gif)
![[F with combining low line]](https://www.rsc.org/images/entities/char_0046_0332.gif)
![[R with combining low line]](https://www.rsc.org/images/entities/char_0052_0332.gif)
![[H with combining low line]](https://www.rsc.org/images/entities/char_0048_0332.gif)
![[R with combining low line]](https://www.rsc.org/images/entities/char_0052_0332.gif)
![[N with combining low line]](https://www.rsc.org/images/entities/char_004e_0332.gif)
![[R with combining low line]](https://www.rsc.org/images/entities/char_0052_0332.gif)
![[K with combining low line]](https://www.rsc.org/images/entities/char_004b_0332.gif)
![[G with combining low line]](https://www.rsc.org/images/entities/char_0047_0332.gif) , OOGCG
, OOGCG![[I with combining low line]](https://www.rsc.org/images/entities/char_0049_0332.gif)
![[K with combining low line]](https://www.rsc.org/images/entities/char_004b_0332.gif)
![[V with combining low line]](https://www.rsc.org/images/entities/char_0056_0332.gif)
![[A with combining low line]](https://www.rsc.org/images/entities/char_0041_0332.gif)
![[V with combining low line]](https://www.rsc.org/images/entities/char_0056_0332.gif) G, CG
G, CG![[R with combining low line]](https://www.rsc.org/images/entities/char_0052_0332.gif)
![[G with combining low line]](https://www.rsc.org/images/entities/char_0047_0332.gif)
![[D with combining low line]](https://www.rsc.org/images/entities/char_0044_0332.gif)
![[S with combining low line]](https://www.rsc.org/images/entities/char_0053_0332.gif) , and CGG
, and CGG![[P with combining low line]](https://www.rsc.org/images/entities/char_0050_0332.gif)
![[H with combining low line]](https://www.rsc.org/images/entities/char_0048_0332.gif)
![[S with combining low line]](https://www.rsc.org/images/entities/char_0053_0332.gif)
![[R with combining low line]](https://www.rsc.org/images/entities/char_0052_0332.gif)
![[N with combining low line]](https://www.rsc.org/images/entities/char_004e_0332.gif) G10
G10![[R with combining low line]](https://www.rsc.org/images/entities/char_0052_0332.gif)
![[G with combining low line]](https://www.rsc.org/images/entities/char_0047_0332.gif)
![[D with combining low line]](https://www.rsc.org/images/entities/char_0044_0332.gif)
![[S with combining low line]](https://www.rsc.org/images/entities/char_0053_0332.gif) ) were synthesized using standard FMOC-chemistry on an automated peptide synthesizer (Liberty Blue™ Automated Microwave Peptide Synthesizer; CEM, Matthews, NC and PS3 Peptide Synthesizer; Protein Technologies, Inc., Tucson, AZ). The peptides were built on Rink Amide MBHA resin (Novabiochem), and all amino acids were double coupled. Peptides were cleaved from resin for 4 hours in 95% triflouroacetic acid (Arcos Organics), 2.5% triisopropylsilane (Arcos Organics), and 2.5% water (all percentages v/v) supplemented with 50 mg mL−1 dithiothreitol (Research Products International). Cleavage solutions for sequences GCRD
) were synthesized using standard FMOC-chemistry on an automated peptide synthesizer (Liberty Blue™ Automated Microwave Peptide Synthesizer; CEM, Matthews, NC and PS3 Peptide Synthesizer; Protein Technologies, Inc., Tucson, AZ). The peptides were built on Rink Amide MBHA resin (Novabiochem), and all amino acids were double coupled. Peptides were cleaved from resin for 4 hours in 95% triflouroacetic acid (Arcos Organics), 2.5% triisopropylsilane (Arcos Organics), and 2.5% water (all percentages v/v) supplemented with 50 mg mL−1 dithiothreitol (Research Products International). Cleavage solutions for sequences GCRD![[V with combining low line]](https://www.rsc.org/images/entities/char_0056_0332.gif)
![[P with combining low line]](https://www.rsc.org/images/entities/char_0050_0332.gif)
![[M with combining low line]](https://www.rsc.org/images/entities/char_004d_0332.gif)
![[S with combining low line]](https://www.rsc.org/images/entities/char_0053_0332.gif) ↓
↓![[M with combining low line]](https://www.rsc.org/images/entities/char_004d_0332.gif)
![[R with combining low line]](https://www.rsc.org/images/entities/char_0052_0332.gif)
![[G with combining low line]](https://www.rsc.org/images/entities/char_0047_0332.gif)
![[G with combining low line]](https://www.rsc.org/images/entities/char_0047_0332.gif) DRCG and CGKG
DRCG and CGKG![[Y with combining low line]](https://www.rsc.org/images/entities/char_0059_0332.gif)
![[I with combining low line]](https://www.rsc.org/images/entities/char_0049_0332.gif)
![[G with combining low line]](https://www.rsc.org/images/entities/char_0047_0332.gif)
![[S with combining low line]](https://www.rsc.org/images/entities/char_0053_0332.gif)
![[R with combining low line]](https://www.rsc.org/images/entities/char_0052_0332.gif) also were supplemented with 25 mg mL−1 phenol (Research Products International). After cleavage, all peptides were precipitated in cold diethyl ether (9× excess volume) overnight at 4 °C and purified by reverse-phase high performance liquid chromatography (HPLC; XBridge BEH C18 OBD 5 μm column; Waters, Milford, MA) with a linear water–acetontrile (ACN) gradient (water
also were supplemented with 25 mg mL−1 phenol (Research Products International). After cleavage, all peptides were precipitated in cold diethyl ether (9× excess volume) overnight at 4 °C and purified by reverse-phase high performance liquid chromatography (HPLC; XBridge BEH C18 OBD 5 μm column; Waters, Milford, MA) with a linear water–acetontrile (ACN) gradient (water![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) ACN 95
ACN 95![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 5 to 45
5 to 45![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 5; 1.17% change in water per minute). Purified peptides were lyophilized, and their molecular weights were verified by mass spectrometry (ESI Fig. S2–S7†).
5; 1.17% change in water per minute). Purified peptides were lyophilized, and their molecular weights were verified by mass spectrometry (ESI Fig. S2–S7†).
2.3. Synthesis and rheological characterization of hydrogels
Monomer stocks were prepared by dissolving each in Dulbecco's phosphate buffer saline (PBS; ThermoFisher; Paisley, Scotland, UK): (i) PEG-8-Nb (150 mM Nb functionality), (ii) lithium phenyl-2,4,6-trimethylbenzoylphospohinate (LAP) (25.5 mM) sterile filtered with a 0.45 μm filter, and (iii) each peptide (30–50 mM). Stock solutions were aliquoted and stored at −80 °C until use. Ellman's assay was performed to determine thiol concentration for each peptide stock prior to freezing.Hydrogel precursor solutions were prepared by diluting stock solutions to a final concentration of 0.72 mM PEG-8-Nb (3 wt%; 5.3 mM Nb functional groups), 1.65 mM peptide crosslinker (3.3 mM SH functional groups), 2 mM LAP, and 2 mM pendant peptide (RGDS, YIGSR, PHSRNG10RGDS, KKQRFRHRNRKG, or IKVAV) in PBS containing 50 U mL−1 penicillin, 50 μg mL−1 streptomycin, and 0.2% fungizone. A PEG-8-Nb concentration of 3 wt% and norbornene![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) thiol stoichiometry of 1
thiol stoichiometry of 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 was used for hydrogel preparation for all cell experiments.
1 was used for hydrogel preparation for all cell experiments.
Rheology measurements were conducted on an AR-G2 rheometer with a UV-visible light attachment (TA instruments, New Castle, DE) in tandem with an Omnicure Series 2000 light source (Excilitas, Waltham, MA) with a 365 nm bandpass filter and light guide (Exfo). Hydrogel precursor solution (10 μL) was placed on the rheometer (8 mm flat plate geometry with a gap of 150 μm), and hydrogel crosslinking and gelation was monitored by measuring storage (G′) and loss (G′′) moduli at 0.5% applied strain and 2 rad s−1 frequency upon irradiation (10 mW cm−2 at 365 nm). Gelation time was based on the change in G′ within 5% between two consecutive points, which was within 2 minutes of irradiation.37 Frequency sweeps at 1% strain were performed after irradiation was complete (2 minutes at 10 mW cm−2) to measure the final moduli of hydrogels formed in situ on the rheometer. All rheometric measurements were performed within the linear viscoelastic regime. Equilibrium moduli were estimated based on previous reports42,43 using this measured modulus and eqn (1), where G0 is the in situ-measured shear modulus; Tfinal is 310 K; T0 is 298 K; and Q0 is the initial volumetric swelling ratio of the hydrogels. Qfinal was calculated using eqn (2), where χ is 0.426, the PEG–water interaction parameter; N is 304, the number of PEG repeats between crosslinks; and ϕ0 is the initial volume fraction of polymer. Lastly, the Young's modulus (E) was estimated by rubber elasticity theory, eqn (3), where ν is Poisson's ratio and is taken to be 0.5 for these elastic PEG hydrogels.
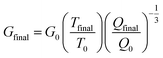 | (1) |
| Qfinal = (1 − 2χ)N0.57ϕ−0.38 | (2) |
| E = 2Gfinal(1 + ν) | (3) |
2.4. Cell culture
IMR90-4 iPSCs were purchased from WiCell® (Madison, WI) and cultured under sterile conditions at 37 °C with 5% CO2 on 6-well tissue culture polystyrene (CELLTREAT Scientific Products, Pepperell, MA) coated with Corning® Matrigel® Basement Membrane Matrix Growth Factor Reduced (Corning; Bedford, MA). IMR90-4 iPSCs are reprogrammed fetal fibroblast cells and have previously shown potential to differentiate into all three germ layers.21 Upon arrival, Matrigel® (5 mL vial) was aliquoted and stored frozen at −80 °C. For plate coating, aliquots (1 mg) were thawed overnight on ice in the refrigerator, and the following day diluted in 12 mL in DMEM/F12 50/50, 1× with L-glutamine, and 15 mM HPEPES media (Corning Cellgro) (83 μg mL−1 Matrigel® in DMEM/F12). The media-Matrigel® solution was applied to plates (1 mL per well) and incubated for 1 hour at room temperature before seeding with 1 × 106 iPSCs per well. From thaw, iPSCs were cultured in growth medium (mTeSR™1 (Stemcell Technologies; Vancouver, BC, Canada) with 10 μM Y-27632 dihydrochloride ROCK Inhibitor (Y-27632) (TOCRIS Bioscience; Bristol, UK)), changing culture medium daily (2 mL per well). Cells were passaged every 4–5 days by dissociation with Versene solution (ThermoFisher Scientific; Grand Island, US). Specifically, iPSCs were incubated at 37 °C for 10 minutes with 1 mL per well of Versene, diluted with mTeSR™1 after dissociation, spun down at 1000 RPM for 5 minutes, resuspended in mTeSR™1, and split 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 4.
4.
2.5. Cell encapsulation
iPSCs were encapsulated as a single cell suspension at a density of 5 × 106 iPSCs per mL in 20 μL of hydrogel precursor solution (100![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 iPSCs per hydrogel), a seeding density in the range of those typically used for cell encapsulations in synthetic hydrogels that provides a balance between cell–cell contact and cell–matrix interactions.44,45 A bulk hydrogel precursor solution as described earlier (section 2.3.) was prepared for the formation of 6 to 9 20 μL hydrogels. iPSCs were dissociated with 1 mL per well of Accutase (Innovative Cell Technologies; San Diego, CA) and rinsed with mTeSR™1. Cells were counted using a hemocytometer, and iPSC aliquots for 6 or 9 gels (at 5 × 106 iPSCs per mL) were spinned down at 1200 RPM for 3 minutes. Cells were re-suspended in mTeSR™1 with 10 μM Y-27632 at the volume required to form gels with a final concentration of PEG-8-Nb, peptide crosslinker, peptide, and LAP of 5.3 mM, 3.3 mM, 2 mM, and 2 mM, respectively, where the cell suspension was used in place of a portion of PBS buffer. Hydrogels were formed in 1 mL syringe molds: 20 μL from the bulk hydrogel prep solution was pipetted into a syringe (end removed), and the hydrogel was formed by irradiating the solution with 10 mW cm−2 at 365 nm for 2 minutes using the Omnicure series 2000 light source (light guide with collimating lens). Three replicates were formed at a time and each placed into a well of a 48-well non-tissue culture treated plate with 300 μL of growth medium unless otherwise noted (e.g., experimental condition of culture without 10 μM Y-27632 or differentiation conditions). Initial culture medium was replaced after 1 hour of incubation to wash away any unreacted monomer from the cells in 3D culture. Cell-gel constructs in culture medium were incubated at 37 °C with 5% CO2 to support cell growth.
000 iPSCs per hydrogel), a seeding density in the range of those typically used for cell encapsulations in synthetic hydrogels that provides a balance between cell–cell contact and cell–matrix interactions.44,45 A bulk hydrogel precursor solution as described earlier (section 2.3.) was prepared for the formation of 6 to 9 20 μL hydrogels. iPSCs were dissociated with 1 mL per well of Accutase (Innovative Cell Technologies; San Diego, CA) and rinsed with mTeSR™1. Cells were counted using a hemocytometer, and iPSC aliquots for 6 or 9 gels (at 5 × 106 iPSCs per mL) were spinned down at 1200 RPM for 3 minutes. Cells were re-suspended in mTeSR™1 with 10 μM Y-27632 at the volume required to form gels with a final concentration of PEG-8-Nb, peptide crosslinker, peptide, and LAP of 5.3 mM, 3.3 mM, 2 mM, and 2 mM, respectively, where the cell suspension was used in place of a portion of PBS buffer. Hydrogels were formed in 1 mL syringe molds: 20 μL from the bulk hydrogel prep solution was pipetted into a syringe (end removed), and the hydrogel was formed by irradiating the solution with 10 mW cm−2 at 365 nm for 2 minutes using the Omnicure series 2000 light source (light guide with collimating lens). Three replicates were formed at a time and each placed into a well of a 48-well non-tissue culture treated plate with 300 μL of growth medium unless otherwise noted (e.g., experimental condition of culture without 10 μM Y-27632 or differentiation conditions). Initial culture medium was replaced after 1 hour of incubation to wash away any unreacted monomer from the cells in 3D culture. Cell-gel constructs in culture medium were incubated at 37 °C with 5% CO2 to support cell growth.
2.6. Assessing cell response to hydrogel formation
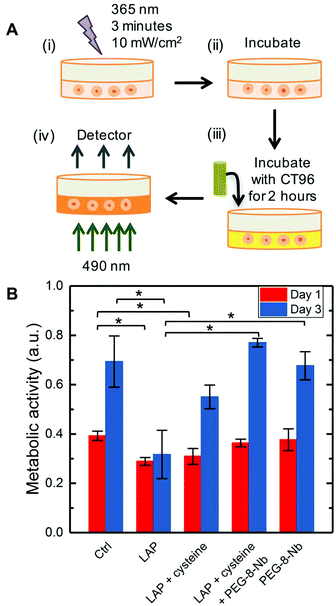
CellTiter96® (CT96) (Promega; Madison, WI), a metabolic activity assay, was used as a measurement of iPSC viability and growth in 2D culture for assessing any impact of the process of hydrogel formation (e.g., monomers, radicals, and light used during photoinitiated step growth polymerization). For this study, iPSCs were dissociated with accutase and seeded (25![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 iPSCs per cm2) on a 96-well Matrigel® coated plate and incubated overnight in 100 μL of growth medium. The next day, different monomer conditions were added to the cells: experimental conditions (n = 3) included LAP, LAP + cysteine, LAP + cysteine + PEG-8-Nb, and PEG-8-Nb each dissolved in PBS and irradiated (10 mW cm−2 at 365 nm for 3 minutes), and a growth medium control. Concentrations of LAP, cysteine, and PEG-8-Nb were the same as those used for iPSC encapsulation at 2 mM, 3.3 mM, and 3 wt%, respectively, with a total volume of 50 μL per well. After irradiation, each well was washed 3× with 50 μL of growth medium, and samples were cultured for up to 3 days, feeding daily (100 μL of growth medium) and assessing metabolic activity on day 1 and day 3. On the day of analysis, 20 μL of CT96 solution was added to each well, incubated with cells for 2 hours (37 °C with 5% CO2), and transferred to a plate reader (BioTek Synergy H4 Hybrid Reader; Winooski, VT, USA) for measurement of absorbance at 490 nm for each condition.
000 iPSCs per cm2) on a 96-well Matrigel® coated plate and incubated overnight in 100 μL of growth medium. The next day, different monomer conditions were added to the cells: experimental conditions (n = 3) included LAP, LAP + cysteine, LAP + cysteine + PEG-8-Nb, and PEG-8-Nb each dissolved in PBS and irradiated (10 mW cm−2 at 365 nm for 3 minutes), and a growth medium control. Concentrations of LAP, cysteine, and PEG-8-Nb were the same as those used for iPSC encapsulation at 2 mM, 3.3 mM, and 3 wt%, respectively, with a total volume of 50 μL per well. After irradiation, each well was washed 3× with 50 μL of growth medium, and samples were cultured for up to 3 days, feeding daily (100 μL of growth medium) and assessing metabolic activity on day 1 and day 3. On the day of analysis, 20 μL of CT96 solution was added to each well, incubated with cells for 2 hours (37 °C with 5% CO2), and transferred to a plate reader (BioTek Synergy H4 Hybrid Reader; Winooski, VT, USA) for measurement of absorbance at 490 nm for each condition.
2.7. Live/Dead viability assay on encapsulated cells
Viability of iPSCs after encapsulation in hydrogels was assessed using a LIVE/DEAD® Viability/Cytotoxicity Kit (ThermoFisher Scientific) on day 1 and day 3. Calcein AM detects esterase activity of cells, producing a fluorescent green dye (ex/em ∼ 495 nm/515 nm) in the cytosol of living cells, whereas ethidium homodimer-1 is a fluorescent red dye (ex/em ∼ 495 nm/635 nm) that binds to nucleic acids and labels the nuclei of cells with damaged membranes indicating dead cells. Briefly, cells were encapsulated in hydrogels as described earlier, and hydrogels (n = 3) were washed 2× with 500 μL of PBS for 5 minutes followed by a 30-minute incubation (37 °C at 5% CO2) with 400 μL of PBS containing calcein AM (2 μM) and ethidium homodimer-1 (4 μM). After staining, hydrogels were again washed (2 × 500 μL of PBS for 5 minutes) before imaging. Hydrogels were transferred to a chamber slide (Nunc Lab-Tek™ II Chamber Slide, Glass, 1 well) and imaged with a confocal microscope (Zeiss LSM 800, 10× objective at a zoom of 0.6× and frame size of 1024 × 1024 for each image, 350 μm z-stack, 3 images per hydrogel sample). Orthogonal projections were made of each z-stack, and live (green) and dead (red) cells were counted using ImageJ. The percentage of viable cells was calculated by the number of green cells/total number of cells × 100%.2.8. Metabolic activity of encapsulated cells
AlamarBlue® cell viability reagent (Thermo Fisher) was used to examine iPSC metabolic activity in hydrogels at days 1, 3, and 7 time points following a modified version of a previously published protocol.44 iPSCs were encapsulated in hydrogels (n = 3) and cultured for up to 7 days. At time points of interest (days 1, 3, and 7), alamarBlue® reagent (10×) was diluted 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 10 in growth medium with 10 μM Y-27632. The culture medium for each hydrogel was replaced with this solution (300 μL per hydrogel cultured in 48-well plate) and incubated for 16 hours (37 °C at 5% CO2). Conditioned culture media were collected from each well, and hydrogels were replenished with fresh standard culture medium. Conditioned media (100 μL from each well) were transferred to a black 96-well plate, and fluorescence was measured (BioTek Synergy H4 Hybrid Reader, ex/em ∼ 560 nm/590 nm).
10 in growth medium with 10 μM Y-27632. The culture medium for each hydrogel was replaced with this solution (300 μL per hydrogel cultured in 48-well plate) and incubated for 16 hours (37 °C at 5% CO2). Conditioned culture media were collected from each well, and hydrogels were replenished with fresh standard culture medium. Conditioned media (100 μL from each well) were transferred to a black 96-well plate, and fluorescence was measured (BioTek Synergy H4 Hybrid Reader, ex/em ∼ 560 nm/590 nm).
2.9. Neural differentiation
iPSCs (5 × 106 iPSCs per mL) were encapsulated in hydrogels presenting YIGSR, PHSRNG10RGDS, or RGDS, as detailed in section 2.6., and cultured in STEMdiff™ Neural Induction Medium (Stem Cell Technologies; Vancouver, Canada) for 6 days for differentiation into NPCs.46,47 After 6 days with daily feedings, encapsulated cells were immunostained (n = 3 per condition), and RNA was isolated for qRT-PCR analysis as detailed below (n = 3 per condition to evaluate pluripotency (oct4) and neural progeny (pax6)). Note, iPSCs were encapsulated in an RGDS-presenting hydrogel and cultured in growth medium as a pluripotent control.2.10. RNA isolation and qRT-PCR
RNA from iPSCs incubated in growth medium or differentiation medium was isolated from plated and hydrogel samples using a Qiagen RNeasy mini kit (Qiagen, Germantown, MD). Specifically, each hydrogel was placed into a microcentrifuge tube and degraded by collagenase (Collagenase, Type II, powder, Thermofisher): 500 μL of 200 U collagenase dissolved in PBS was added per hydrogel, and tubes were incubated at 37 °C at 5% CO2 for 40–60 minutes until the hydrogels were observed to be completely degraded (e.g., a liquid solution that could be freely pipetted). Tubes were centrifuged at 1200 RPM for 3 minutes to collect cells, and cells were washed (1200 RPM for 3 minutes) once with 500 μL of PBS to remove any residual collagenase and PEG. Cells isolated from two hydrogel samples were combined and lysed for 5 minutes in 600 μL of lysis buffer for each RNA isolation sample. This approach resulted in RNA of a sufficient quantity and solution of appropriate volume for subsequent processing. Each lysis buffer solution (n = 3) subsequently was placed into a spin column for RNA purification. All samples were DNase treated for 15 minutes at room temperature, and 50 μL of RNase-free water was used to collect RNA. The concentration of RNA isolated from each sample was assessed by a Nanodrop 2000c spectrophotometer (Thermofisher; Waltham, MA) and observed to be 20–200 ng μL−1 with a A260/A280 > 2.0 (n = 3 RNA isolations per culture condition).After RNA isolation, an iTaq™ Universal One-Step RT-qPCR kit (Bio-Rad; Hercules, CA) was used for qRT-PCR analysis. oct4 and pax6 gene expression were measured to assess pluripotency and neural progeny, respectively, and GAPDH was used as the housekeeping gene for analysis. A total reaction mix volume of 15 μL was run for each sample on a CFX96 Touch™ Real-Time PCR Detection System (Bio-Rad; Hercules, CA) to measure Ct values. These reactions included 300 nM forward and reverse primers (primers listed in ESI Table 1†), 0.188 μL iScript reverse transcriptase, 7.5 μL iTaq universal SYBR® Green reaction mix (2×), 100 ng of RNA, and nuclease-free H2O for the remaining volume. ΔCt values were calculated by subtracting the housekeeping Ct value from the gene of interest (oct4 or pax6) Ct value for each condition after differentiation. ΔΔCt values were calculated as the difference between ΔCt's of the experimental condition after differentiation and a 3D pluripotent control (iPSCs in RGD-presenting hydrogels cultured in growth medium). An example analysis for calculating ΔCt and ΔΔCt is shown in eqn (4)–(7) (example, iPSCs in YIGSR-presenting hydrogels cultured in differentiation medium analyzed for oct4 expression).
| ΔCtiPSC,oct4 = Ctoct4,iPSC − CtGapdh,iPSC | (4) |
| ΔCtYIGSR,oct4 = Ctoct4,YIGSR − CtGapdh,YIGSR | (5) |
| ΔΔCtYIGSR,oct4 = ΔCtYIGSR,oct4 − ΔCtiPSC,oct4 | (6) |
| Relative expresion = 2−ΔΔCtYIGSR,oct4 | (7) |
A two-sided Mann–Whitney test was used to determine statistical significance between samples. Three biological replicates with 2 technical replicates were used for each culture condition.
2.11. Immunocytochemistry
Encapsulated iPSCs incubated in growth medium or differentiation medium were immunostained for OCT4 and PAX6, markers of pluripotency and NPCs, respectively. At time points of interest, culture media were removed, and hydrogels were rinsed (2× with 300 μL of PBS for 5 minutes). Cells were fixed with cold 4% PFA (300 μL) at 4 °C for 15 minutes and rinsed (1× with 300 μL of PBS, 2× with 300 μL PBS containing 1% BSA and 0.1% TritonX). Cells were then permeabilized and blocked with 1% BSA and 0.1% TritonX in PBS for 45 minutes. Primary antibodies (mouse anti-OCT4 (Santa Cruz Biotechnology, CA) and rabbit anti-PAX6 (ThermoFisher)) subsequently were added at 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 250 dilution in PBS with 1% BSA and 0.1% TritonX and incubated at 4 °C overnight. The next day, hydrogels were washed (3× in PBS with 1% BSA and 0.1% TritonX) for 45 minutes. Secondary antibodies AF488 goat anti-mouse IgG (H + L) (ThermoFisher Scientific; Eugene, OR) and AF647 goat anti-rabbit IgG (H + L) (ThermoFisher Scientific; Eugene, OR) were added at a dilution of 1
250 dilution in PBS with 1% BSA and 0.1% TritonX and incubated at 4 °C overnight. The next day, hydrogels were washed (3× in PBS with 1% BSA and 0.1% TritonX) for 45 minutes. Secondary antibodies AF488 goat anti-mouse IgG (H + L) (ThermoFisher Scientific; Eugene, OR) and AF647 goat anti-rabbit IgG (H + L) (ThermoFisher Scientific; Eugene, OR) were added at a dilution of 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 250 in PBS with 1% BSA and 0.1% TritonX and incubated at 4 °C overnight. The next day, hydrogels were washed (3× in PBS with 1% BSA and 0.1% TritonX for 30 minutes each). Before imaging, samples were stained with DAPI in PBS (700 nM) for 30 minutes at room temperature and washed (3× with PBS for 30 minutes). Samples were imaged with confocal microscopy (Zeiss LSM800, 20× objective, Z-stacks of 50–100 μm, frame size of 1593 × 1593 pixels). Z-stack images were orthogonally projected for ease of viewing and analysis, and the area of cell clusters was measured using Zen (blue edition, Zeiss).
250 in PBS with 1% BSA and 0.1% TritonX and incubated at 4 °C overnight. The next day, hydrogels were washed (3× in PBS with 1% BSA and 0.1% TritonX for 30 minutes each). Before imaging, samples were stained with DAPI in PBS (700 nM) for 30 minutes at room temperature and washed (3× with PBS for 30 minutes). Samples were imaged with confocal microscopy (Zeiss LSM800, 20× objective, Z-stacks of 50–100 μm, frame size of 1593 × 1593 pixels). Z-stack images were orthogonally projected for ease of viewing and analysis, and the area of cell clusters was measured using Zen (blue edition, Zeiss).
2.12. Statistical analysis
All values are reported as mean ± standard error (n = 3 for each condition). Unless otherwise noted, statistical significance was determined by a two-sided Student T-test.3. Results and discussion
3.1. Design and formation of hydrogels
We aimed to establish an approach for the encapsulation and culture of iPSCs within photoinitiated PEG-peptide-based synthetic matrices and to utilize this system to test hypotheses about key cell–matrix interactions for both maintaining iPSC viability and permitting their differentiation. We selected the rapid photoinitiated thiol–norbornene reaction for encapsulation of iPSCs owing to its successful use with a variety of other cell types, including primary human mesenchymal stem cells and neural cells derived from ESCs.37,42 Here, a PEG-8-Nb macromer was crosslinked with a MMP-degradable sequence GCRD![[V with combining low line]](https://www.rsc.org/images/entities/char_0056_0332.gif)
![[P with combining low line]](https://www.rsc.org/images/entities/char_0050_0332.gif)
![[M with combining low line]](https://www.rsc.org/images/entities/char_004d_0332.gif)
![[S with combining low line]](https://www.rsc.org/images/entities/char_0053_0332.gif) ↓
↓![[M with combining low line]](https://www.rsc.org/images/entities/char_004d_0332.gif)
![[R with combining low line]](https://www.rsc.org/images/entities/char_0052_0332.gif)
![[G with combining low line]](https://www.rsc.org/images/entities/char_0047_0332.gif)
![[G with combining low line]](https://www.rsc.org/images/entities/char_0047_0332.gif) DRCG, which is responsive to MMP-2 amongst other cell-secreted enzymes that are known to be upregulated in iPSCs allowing cell-driven remodeling of the resulting synthetic matrix.20 Additionally, pendant peptides (e.g., CG
DRCG, which is responsive to MMP-2 amongst other cell-secreted enzymes that are known to be upregulated in iPSCs allowing cell-driven remodeling of the resulting synthetic matrix.20 Additionally, pendant peptides (e.g., CG![[R with combining low line]](https://www.rsc.org/images/entities/char_0052_0332.gif)
![[G with combining low line]](https://www.rsc.org/images/entities/char_0047_0332.gif)
![[D with combining low line]](https://www.rsc.org/images/entities/char_0044_0332.gif)
![[S with combining low line]](https://www.rsc.org/images/entities/char_0053_0332.gif) ) facilely were incorporated during hydrogel formation by inclusion of a cysteine within the sequence to promote the binding of specific receptors on iPSCs, as detailed below. The hydrogel was rapidly formed by light-triggered thiol–ene ‘click’ chemistry using the photoinitiator LAP, as thiols presented by cysteines on the crosslinker or pendant peptides react in a step-wise fashion with the alkene group on PEG-8-Nb (Fig. 1A).48 Rheological measurements were performed on the hydrogels to determine the gelation time and modulus for this system. The crossover point, where storage modulus equals the loss modulus (G′ = G′′), correlates with the gel point and was observed at ∼t = 1.52 ± 0.07 minutes (Fig. 1C), indicating rapid, light-triggered hydrogel formation. The modulus of the resulting hydrogels was varied with different PEG-8-Nb concentrations in the gel-forming solutions (3 to 6 wt% PEG-8-Nb), where a range of moduli relevant for mimicking a variety of soft tissues was achieved (Young's modulus (E) ∼ 3–13 kPa) (Fig. 1D). In particular, the 3 wt% PEG-8-Nb concentration was selected for use in all following experiments with E ∼ 3 kPa, which is on the same order of magnitude as the modulus of developing neural tissue during embryogenesis (E ∼ 1 kPa)49 and resulted in robust and consistent hydrogel formation.
) facilely were incorporated during hydrogel formation by inclusion of a cysteine within the sequence to promote the binding of specific receptors on iPSCs, as detailed below. The hydrogel was rapidly formed by light-triggered thiol–ene ‘click’ chemistry using the photoinitiator LAP, as thiols presented by cysteines on the crosslinker or pendant peptides react in a step-wise fashion with the alkene group on PEG-8-Nb (Fig. 1A).48 Rheological measurements were performed on the hydrogels to determine the gelation time and modulus for this system. The crossover point, where storage modulus equals the loss modulus (G′ = G′′), correlates with the gel point and was observed at ∼t = 1.52 ± 0.07 minutes (Fig. 1C), indicating rapid, light-triggered hydrogel formation. The modulus of the resulting hydrogels was varied with different PEG-8-Nb concentrations in the gel-forming solutions (3 to 6 wt% PEG-8-Nb), where a range of moduli relevant for mimicking a variety of soft tissues was achieved (Young's modulus (E) ∼ 3–13 kPa) (Fig. 1D). In particular, the 3 wt% PEG-8-Nb concentration was selected for use in all following experiments with E ∼ 3 kPa, which is on the same order of magnitude as the modulus of developing neural tissue during embryogenesis (E ∼ 1 kPa)49 and resulted in robust and consistent hydrogel formation.
3.2. Mechanism of hydrogel formation has limited impact on iPSC metabolic activity
Photoinitiated polymerizations have been used for the encapsulation of a variety of cell types, including primary cells such as human mesenchymal stem cells, hepatic stellate cells, and pancreatic beta cells.42,48,50–52 However, this reaction mechanism for hydrogel formation generates and utilizes free-radicals, which have the potential to be cytotoxic and cause cell death. To examine any potential cytotoxic effects of photopolymerization conditions on iPSCs, iPSC metabolic activity in 2D culture was examined in the presence of the different combinations of precursors and light, as typically used for hydrogel formation: the photoinitiator LAP; LAP + cysteine that is the thiol source in the crosslinker and pendant peptides; LAP + cysteine + PEG-8-Nb macromer; or PEG-8-Nb. iPSCs were seeded onto a Matrigel coated 96-well plate and incubated with these different combinations of hydrogel precursors in growth medium. Light was applied as is done for hydrogel formation (10 mW cm−2 at 365 nm for 3 minutes) followed by rinses with fresh culture medium. Metabolic activity was measured at 1 and 3 days after exposure in comparison to the control (cells cultured in growth medium) to assess any immediate and continued impact of hydrogel formation conditions on cell function (Fig. 2A). As expected, a statistically significant decrease in metabolic activity was observed after exposure to LAP and LAP + cysteine with light, where there is nothing to ‘consume’ the free radicals generated, as compared to cells cultured in the growth medium control (Fig. 2B). Importantly, when macromer was introduced for reaction with the generated thiyl radicals (LAP + cysteine + PEG-8-Nb) or by itself (PEG-8-Nb) and light was applied, no significant changes in iPSC metabolic activity were observed as compared to the control. These data supported the hypothesis that, with all hydrogel-forming components present, iPSC viability would not be significantly impacted during the process of cell encapsulation. Further, 3 days after exposure, all conditions tested except for LAP with light exhibited no statistical difference compared to the control, supporting recovery of cells after an initial insult of thiyl radicals generated by LAP + cysteine but not from the radicals generated by LAP alone. Taken together, these data suggest that generation of free radicals was cytotoxic to iPSCs, but with monomers present, free radicals were sufficiently consumed during step-growth polymerization to appropriately limit impact on cells.3.3. Presentation of an integrin binding peptide and inclusion of ROCK inhibitor improved iPSC viability in 3D culture within PEG-peptide hydrogels initially and over time
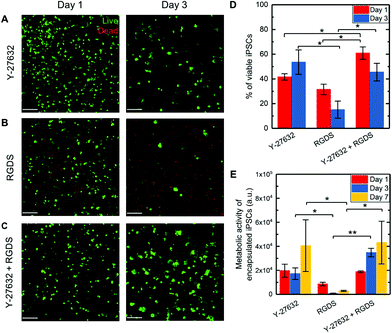
ROCK inhibitor Y-27632 is known to improve iPSC viability during their culture as single cells on Matrigel or within polymeric hydrogels (PNIPAAm-PEG) by both preventing apoptosis and promoting cell–matrix interactions, as noted in the Introduction and discussed further below.16,28 Exploiting the control of matrix composition afforded by PEG-peptide hydrogels, we wanted to test if integrin binding peptides alone or in conjunction with ROCK inhibition could support the viability of iPSCs encapsulated as single cells in a well-defined 3D environment. iPSCs first were cultured in PEG hydrogels without integrin-binding sequences or a ROCK inhibitor (‘blank’ control) and exhibited poor viability (15 ± 2% and 12 ± 2% at days 1 and 3, ESI Fig. S8†), as expected based on prior work in the literature.16 We hypothesized that the presentation of peptides that promote integrin-binding and cell adhesion and addition of a ROCK inhibitor would enhance the viability of these cells. To test this, we examined encapsulated iPSC viability in response to the following conditions: encapsulation in hydrogels cultured with Y-27632, hydrogels presenting covalently-linked RGDS to promote cell adhesion to the matrix, or hydrogels presenting RGDS and cultured with Y-27632. iPSCs in 3D culture subsequently were stained with a Live/Dead membrane integrity assay on days 1 and 3 after encapsulation for all three conditions (Fig. 3A–C). One day after encapsulation, iPSCs within hydrogels presenting RGDS and cultured with Y-27632 had the highest viability of 61 ± 5%, a statistically significant difference compared to RGDS only (32 ± 4%) or Y-27632 only (42 ± 3%) (Fig. 3D). No statistical difference in viability was observed between the RGDS only or Y-27632 only condition at this early time point, whereas both conditions were statistically higher than the ‘blank’ control hydrogel (p < 0.05). These observations at early times after cell encapsulation suggest that presentation of RGDS, which is known to bind several different integrins and promotes general cell adhesion to the matrix,38 has a similar impact on iPSC viability to ROCK inhibition, which is known to prevent apoptosis and increase integrin expression.16,28 At three days of culture, iPSCs encapsulated in hydrogels supplemented with Y-27632 with or without RGDS peptide had the highest viability of 46 ± 7% and 54 ± 10%, respectively. Both cultures supplemented with Y-27632 had statistically higher viability than either the ‘blank’ control hydrogel (12 ± 2%, p < 0.05, ESI Fig. S8†) or the hydrogel with RGDS only (15 ± 7%, p < 0.05, Fig. 3D). These observations at later times after encapsulation support that culture with ROCK inhibitor Y-27632 was important for cell survival in this PEG-peptide hydrogel with either non-specific or no biochemical cues. Similar trends in viability were observed with measurements of metabolic activity over 1 week (Fig. 3E). After 3 days in culture, a statistically significant increase in viability was observed for cells in the RGDS and Y-27632 condition (p < 0.01), compared to iPSCs cultured with RGDS only that exhibited low metabolic activity. Additionally, over 7 days of culture, a trend of increasing metabolic activity was observed for Y-27632 cultures with and without RGDS. Taken together, these data support the relevance of including both integrin binding sequences and ROCK inhibitor for iPSC survival as a single cell suspension in these 3D cultures at both early and late times after encapsulation.Note, ROCK plays a role in actin-myosin contraction, where phosphorylation of myosin light chain (MLC) by ROCK activates myosin II heavy chain-actin contraction, which is known to reduce viability of dissociated pluripotent stem cells.53 Treatment with ROCK inhibitor Y-27632 prevents phosphorylation of MLC, inhibiting myosin-actin contraction and likely thereby improving viability. Although prevention of myosin-actin contraction is not directly tested here, we observed increased viability of iPSCs encapsulated as single cells in 3D hydrogel culture in the presence of Y-27632. Our findings are consistent with other studies in hydrogels permissive to iPSC growth: for example, Lei et al. utilized ROCK inhibitor for culture of iPSCs starting from a single cell suspension in thermoreversible PNIPAAm-PEG hydrogels over 4 days.16 Here, we observed increased viability of iPSCs for 7 days in culture with Y-27632 in a photopolymerized, cell-degradable PEG-peptide hydrogel. From this analysis, ROCK inhibitor Y-27632 was used in all subsequent culture experiments while the more general cell adhesive peptide RGDS was replaced with sequences with more specific binding of integrins of interest. Further, in addition to ROCK inhibition, we examined if increasing cell seeding density (106, 5 × 106, and 107 cells per mL), toward increasing cell–cell interactions at early times in 3D culture, could improve iPSC viability (ESI Fig. S9†); however, no difference in iPSC viability was observed amongst these conditions. Consequently, the intermediate cell seeding density of 5 × 106 iPSCs per mL was used for all other encapsulations.
3.4. Bioinspired integrin binding peptides promoted iPSC viability in 3D culture
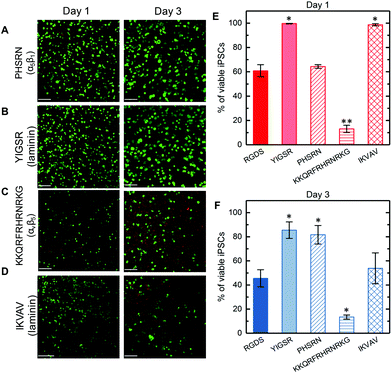
As noted in the Introduction, integrin subunits α5, α6, αv, β1, and β5 have prominent expression on iPSCs and promote their adhesion and proliferation on Matrigel.26 Synthetic PEG hydrogels provide a blank-slate matrix to test various peptide sequences for assessing their impact on iPSCs in 3D culture, where peptides of interest can be facilely incorporated by click chemistry. We hypothesized that peptides associated with binding of these key integrins present on iPSCs could rescue cell viability in 3D culture within such synthetic matrices (Fig. 4A–D). Peptide sequences tested included multi-integrin binding peptide RGDS, laminin-derived YIGSR (from β1 chain) and IKVAV (from α1 chain), PHSRNG10RGDS that binds to α5β1, and KKQRFRHRNRKG that binds to αvβ5 (Fig. 1B).54–56 After 1 day of culture, cells cultured in hydrogels presenting IKVAV or YIGSR exhibited nearly 100% viability (Fig. 4E), which was statistically higher than with the multi-integrin binding RGDS (61 ± 5%) or PHSRNG10RGDS (64 ± 2%). Additionally, iPSCs cultured within hydrogels presenting the αvβ5-binding peptide KKQRFRHRNRKG had the lowest viability (13 ± 3%), statistically lower than iPSCs cultured in RGDS-presenting hydrogels, and continued cell death was observed at day 3 (Fig. 4F). After 3 days of culture, iPSCs cultured in hydrogels presenting PHSRNG10RGDS and YIGSR had the highest viability of 82 ± 8% and 85 ± 7%, respectively, whereas those in IKVAV condition had decreased viability of 54 ± 13%. Similar trends were observed for iPSC metabolic activity, although with increased variability (ESI Fig. S10†).Our approach, inspired by iPSC binding and integrin expression on Matrigel, identified YIGSR or PHSRNG10RGDS as key peptide sequences in PEG-peptide hydrogels for supporting the viability of single cell iPSCs when cultured in three-dimensions over the course of 1 week. Notably, YIGSR and PHSRNG10RGDS peptides both are associated with the binding of β1 integrin, suggesting the potential importance of this integrin for iPSC survival in such 3D cultures. Interestingly, iPSCs cultured within hydrogels presenting the αvβ5-binding peptide KKQRFRHRNRKG exhibited the lowest viability in 3D culture. Although the αvβ5 integrin is associated with iPSC viability for 2D culture on Matrigel, these data suggest that either this heterodimer integrin or the selected integrin-binding peptide sequence alone were not sufficient for promoting iPSC viability in 3D culture.8 Additionally, iPSCs cultured in hydrogels presenting IKVAV that is derived from laminin, one of the main ECM proteins found in Matrigel,57 exhibited intermediate viability in 3D culture. Taken together, the base synthetic matrix established here may prove useful for future investigations in 3D culture of these and other peptides individually or in potential synergistic combinations for promoting iPSC viability, growth, or differentiation.
3.5. β1 integrin is important for iPSC viability in 3D culture within PEG-peptide hydrogels
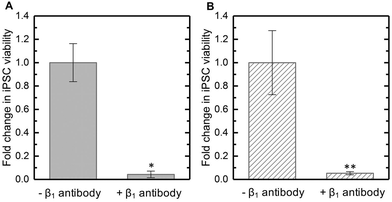
Hydrogels presenting either peptide YIGSR or PHSRNG10RGDS supported the highest iPSC viability during 3D culture in PEG-peptide hydrogels. Since both peptides are associated with binding to β1 integrin, we hypothesized that this integrin may be key for iPSC survival and growth in these 3D cultures. To test this, we applied an anti-β1 antibody, which is widely used in the literature to block β1 integrin,19,58 to iPSCs prior to encapsulation and throughout 3D culture. Viability of iPSCs cultured with the β1-blocking antibody in both hydrogels presenting YIGSR or PHSRNG10RGDS was decimated as compared to respective control hydrogels containing iPSCs cultured without the β1-blocking antibody by day 3 in culture (Fig. 5A and B). Blocking of β1 integrin was confirmed by immunostaining for β1: cells cultured without the β1 antibody stained positive, indicating the presence and availability of β1 integrin, whereas cells cultured with the β1 antibody did not stain positive, further supporting successful blocking (ESI Fig. S11†).This study demonstrated that β1 integrin was key for iPSC survival during 3D culture within these well-defined synthetic matrices. Note, the viability of encapsulated iPSCs with β1 integrin blocked was significantly lower than that observed with no integrin-binding peptide (Fig. 3D). Blocking of β1 integrin will prevent iPSC interactions with relevant ligands presented by peptides covalently attached to the synthetic matrix (here, YIGSR or PHSRNG10RGDS) or ECM proteins potentially present in culture medium or secreted locally by the cells. In light of the observed impact of blocking on cell viability in these 3D cultures, we speculate that blocking of β1 integrin may mitigate some of the beneficial effects of ROCK inhibition, which include increasing integrin expression.28 The materials and approaches established here provide opportunities for future examination of these complex interactions toward insights into the interplay between β1 integrin and ROCK.
3.6. PEG-peptide synthetic matrices are permissive to the growth and differentiation of iPSCs into NPCs
After establishing a well-defined hydrogel matrix that maintained iPSC viability, we wanted to confirm that these matrices were permissive to the differentiation of iPSCs into NPCs. NPCs, amongst the variety of progenitor cell lineages that can be generated by differentiation of iPSCs, are of importance because they are precursor cells to all neural cells: these neural lineages include neurons, astrocytes, and oligodendrocytes that are of interest for both neurodegenerative-related studies and regenerative medicine applications.59,60 We examined differentiation in three conditions: hydrogels presenting RGDS, which generally promotes adhesion, and YIGSR and PHSRNG10RGDS, which promoted iPSC viability in 3D culture. Encapsulated iPSCs in each of these conditions were differentiated for 6 days, using an established differentiation medium that contains key soluble factors (STEMdiff neural induction medium), and compared to 6 days of culture in hydrogels presenting RGDS, YIGSR, and PHSRNG10RGDS and cultured in growth medium (pluripotent control). As expected, iPSCs cultured within RGDS-, YIGSR-, and PHSRNG10RGDS-presenting hydrogels in growth medium formed clusters of cells that were OCT4+, indicating the cells remained pluripotent during culture (Fig. 6A and ESI Fig. S12†). After culture in differentiation medium with hydrogels presenting RGDS, YIGSR, or PHSRNG10RGDS, cells again formed clusters but exhibited a loss of OCT4 expression and gain of PAX6+ expression, a neural progenitor cell marker, indicating differentiation (Fig. 6B–D). To confirm this, qRT-PCR was performed to examine gene expression of the cells cultured in differentiation medium relative to the pluripotent control (Fig. 6E and F). Oct4 expression decreased at least 4-fold for the three matrices cultured in differentiation medium and was statistically lower than the control. Cells in the YIGSR condition had the least oct4 downregulation after differentiation, statistically higher than cells in the RGDS and PHSRNG10RGDS conditions (p < 0.05). Pax6 expression overall increased about 10-fold for all differentiation conditions and was statistically higher than the control (p < 0.01).For all three matrix compositions, we observed an overall decrease in pluripotency (oct4) and increase in neural progeny (pax6) at both the gene and protein levels when cultured in differentiation medium. While oct4 expression is downregulated in all differentiation conditions, the RGDS and PHSRNG10RGDS cultures were most significantly downregulated in comparison to the YIGSR condition. Interestingly, iPSCs in YIGSR-presenting hydrogels had the highest viability amongst all conditions on 1 and 3 days and had the highest metabolic activity over the course of the week; in this context, we speculate that this support of iPSC viability and growth may influence iPSC pluripotency, causing oct4 to be least downregulated even after differentiation. However, with culture in differentiation medium, all hydrogel compositions were observed to support or promote similar expression of the neural progenitor marker (pax6) at the gene level.
In addition to changes in gene and protein expression, differences in cell cluster size were observed after 6 days of differentiation in the three hydrogel compositions (Fig. 6G and ESI Fig. S13†). The average cell cluster area after differentiation was 4000 ± 1000 μm2, 16![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 ± 2000 μm2, or 33
000 ± 2000 μm2, or 33![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 ± 5000 μm2 for hydrogels presenting RGDS, YIGSR, or PHSRNG10RGDS, respectively (p < 0.01). These clusters may have resulted from iPSC migration toward each other to form clusters or enhanced their proliferation in specific matrix compositions; although not tested, this observation may indicate that factors such as cell–cell contact in addition to cell-matrix interactions are important during iPSC growth and neural differentiation in 3D culture environments. Indeed, cell–cell contact (E-cadherin) has been shown to be an important regulator of iPSC fate in 2D culture, where cluster size was controlled by confinement to micropatterned adhesive regions (fibronectin) of different diameter or geometry.61 Additionally, cell–cell contact has been shown to promote NPC progeny in 3D culture in highly-degradable hydrogel-based matrices, as cells degrade and remodel the matrix.45 Light-based chemistries such as the photoinitiated thiol–ene reaction used in this work provide opportunities for future studies that probe the role of cell–cell and cell–matrix interactions in iPSC function and fate in three dimensions. For example, the capability of photopolymerization to generate patterns of peptides in multiple dimensions could be exploited to examine the individual and combinatorial of effects of these interactions on pluripotency or differentiation (e.g., effects of RGDS vs. PHSRN vs. PHSRNG10RGDS or PHSRNG10RGDS vs. cadherin mimics such as HAV)56,62,63 or direct downstream cellular functions (e.g., process extension of terminally-differentiated neuronal cells).64,65
000 ± 5000 μm2 for hydrogels presenting RGDS, YIGSR, or PHSRNG10RGDS, respectively (p < 0.01). These clusters may have resulted from iPSC migration toward each other to form clusters or enhanced their proliferation in specific matrix compositions; although not tested, this observation may indicate that factors such as cell–cell contact in addition to cell-matrix interactions are important during iPSC growth and neural differentiation in 3D culture environments. Indeed, cell–cell contact (E-cadherin) has been shown to be an important regulator of iPSC fate in 2D culture, where cluster size was controlled by confinement to micropatterned adhesive regions (fibronectin) of different diameter or geometry.61 Additionally, cell–cell contact has been shown to promote NPC progeny in 3D culture in highly-degradable hydrogel-based matrices, as cells degrade and remodel the matrix.45 Light-based chemistries such as the photoinitiated thiol–ene reaction used in this work provide opportunities for future studies that probe the role of cell–cell and cell–matrix interactions in iPSC function and fate in three dimensions. For example, the capability of photopolymerization to generate patterns of peptides in multiple dimensions could be exploited to examine the individual and combinatorial of effects of these interactions on pluripotency or differentiation (e.g., effects of RGDS vs. PHSRN vs. PHSRNG10RGDS or PHSRNG10RGDS vs. cadherin mimics such as HAV)56,62,63 or direct downstream cellular functions (e.g., process extension of terminally-differentiated neuronal cells).64,65
4. Conclusions
We have established well-defined, photopolymerizable synthetic matrices that enable the encapsulation, culture, and differentiation of iPSCs in 3D culture. Initial examinations of metabolic activity of iPSCs demonstrated the compatibility of the selected photopolymerization conditions for iPSC culture. ROCK inhibitor was confirmed as important for single-cell suspension culture of iPSCs in three dimensions over time. With the base matrix and culture conditions established, ligands and protein-derived peptides inspired by iPSC interactions with Matrigel were identified to further enhance cell viability, specifically YIGSR and PHSRNG10RGDS, where β1 was observed to be key. Lastly, these matrices were determined to be permissive to iPSC differentiation into NPCs. Interestingly, cluster size of the resulting NPCs varied amongst the different peptide compositions, with the largest cluster size in PHSRNG10RGDS gels. This observation may indicate that other properties, such as cell–cell contact, play a role in iPSC differentiation or function within these materials for future investigations. In sum, we have established a synthetic 3D culture system for the encapsulation and culture of iPSCs with relevance for examining key microenvironment properties that regulate iPSC function and fate in three dimensions and potential utility for 3D models in the study of disease.Conflicts of interest
The authors have no conflict of interests to report.Acknowledgements
The authors would like to acknowledge support, for related work in their laboratories, from the Delaware COBRE programs with grants from the National Institute of General Medicine Sciences (NIGMS P20GM104316 and 5 P30 GM110758-02) from the National Institutes of Health, the Susan G. Komen Foundation (CCR16377327), the PEW Charitable Trusts (0026178), the Burroughs Wellcome Fund (1006787), and the University of Delaware Research Foundation.References
- K. Takahashi, K. Tanabe, M. Ohnuki, M. Narita, T. Ichisaka, K. Tomoda and S. Yamanaka, Cell, 2007, 131, 861–872 CrossRef CAS PubMed.
- V. Tabar and L. Studer, Nat. Rev. Genet., 2014, 15, 82–92 CrossRef CAS PubMed.
- Z. Jiang, Y. Han and X. Cao, Cell. Mol. Immunol., 2014, 11, 17–24 CrossRef CAS PubMed.
- D. Paull, A. Sevilla, H. Zhou, A. K. Hahn, H. Kim, C. Napolitano, A. Tsankov, L. Shang, K. Krumholz, P. Jagadeesan, C. M. Woodard, B. Sun, T. Vilboux, M. Zimmer, E. Forero, D. N. Moroziewicz, H. Martinez, M. C. V. Malicdan, K. A. Weiss, L. B. Vensand, C. R. Dusenberry, H. Polus, K. T. L. Sy, D. J. Kahler, W. A. Gahl, S. L. Solomon, S. Chang, A. Meissner, K. Eggan and S. A. Noggle, Nat. Methods, 2015, 12, 885–892 CrossRef CAS PubMed.
- V. Akopian, P. W. Andrews, S. Beil, N. Benvenisty, J. Brehm, M. Christie, A. Ford, V. Fox, P. J. Gokhale, L. Healy, F. Holm, O. Hovatta, B. B. Knowles, T. E. Ludwig, R. D. G. McKay, T. Miyazaki, N. Nakatsuji, S. K. W. Oh, M. F. Pera, J. Rossant, G. N. Stacey and H. Suemori, In Vitro Cell. Dev. Biol.: Anim., 2010, 46, 247–258 CrossRef PubMed.
- G. Chen, D. R. Gulbranson, Z. Hou, J. M. Bolin, V. Ruotti, M. D. Probasco, K. Smuga-Otto, S. E. Howden, N. R. Diol, N. E. Propson, R. Wagner, G. O. Lee, J. Antosiewicz-Bourget, J. M. C. Teng and J. A. Thomson, Nat. Methods, 2011, 8, 424–429 CrossRef CAS PubMed.
- S. Rodin, L. Antonsson, C. Niaudet, O. E. Simonson, E. Salmela, E. M. Hansson, A. Domogatskaya, Z. Xiao, P. Damdimopoulou, M. Sheikhi, J. Inzunza, A.-S. Nilsson, D. Baker, R. Kuiper, Y. Sun, E. Blennow, M. Nordenskjöld, K.-H. Grinnemo, J. Kere, C. Betsholtz, O. Hovatta and K. Tryggvason, Nat. Commun., 2014, 5, 3195 Search PubMed.
- T. Miyazaki, S. Futaki, H. Suemori, Y. Taniguchi, M. Yamada, M. Kawasaki, M. Hayashi, H. Kumagai, N. Nakatsuji, K. Sekiguchi and E. Kawase, Nat. Commun., 2012, 3, 1236 CrossRef PubMed.
- S. Jin, H. Yao, J. L. Weber, Z. K. Melkoumian and K. Ye, PLoS One, 2012, 7, e50880 CAS.
- S. R. Braam, L. Zeinstra, S. Litjens, D. Ward-van Oostwaard, S. van den Brink, L. van Laake, F. Lebrin, P. Kats, R. Hochstenbach, R. Passier, A. Sonnenberg and C. L. Mummery, Stem Cells, 2008, 26, 2257–2265 CrossRef CAS PubMed.
- J. R. Klim, L. Li, P. J. Wrighton, M. S. Piekarczyk and L. L. Kiessling, Nat. Methods, 2010, 7, 989–994 CrossRef CAS PubMed.
- Y. Wang, L. Cheng and S. Gerecht, Ann. Biomed. Eng., 2014, 42, 1357–1372 CrossRef CAS PubMed.
- C. Chung, B. L. Pruitt and S. C. Heilshorn, Biomater. Sci., 2013, 1, 1082 RSC.
- L. Schukur, P. Zorlutuna, J. M. Cha, H. Bae and A. Khademhosseini, Adv. Healthcare Mater., 2013, 2, 195–205 CrossRef CAS PubMed.
- A. M. Hilderbrand, E. M. Ovadia, M. S. Rehmann, P. M. Kharkar, C. Guo and A. M. Kloxin, Curr. Opin. Solid State Mater. Sci., 2016, 20, 213–219 CrossRef PubMed.
- Y. Lei and D. V. Schaffer, Proc. Natl. Acad. Sci. U. S. A., 2013, 110, E5039–E5048 CrossRef CAS PubMed.
- S. Gerecht, J. A. Burdick, L. S. Ferreira, S. A. Townsend, R. Langer and G. Vunjak-Novakovic, Proc. Natl. Acad. Sci. U. S. A., 2007, 104, 11298–11303 CrossRef PubMed.
- A. Ranga, S. Gobaa, Y. Okawa, K. Mosiewicz, A. Negro and M. P. Lutolf, Nat. Commun., 2014, 5, 4324 CAS.
- S. T. Lee, J. I. Yun, Y. S. Jo, M. Mochizuki, A. J. van der Vlies, S. Kontos, J. E. Ihm, J. M. Lim and J. A. Hubbell, Biomaterials, 2010, 31, 1219–1226 CrossRef CAS PubMed.
- M. Jang, S. T. Lee, J. W. Kim, J. H. Yang, J. K. Yoon, J. C. Park, H. M. Ryoo, A. J. van der Vlies, J. Y. Ahn, J. A. Hubbell, Y. S. Song, G. Lee and J. M. Lim, Biomaterials, 2013, 34, 3571–3580 CrossRef CAS PubMed.
- J. Yu, M. Vodyanik and K. Smuga-Otto, Science, 2007, 318, 1917–1920 CrossRef CAS PubMed.
- I.-H. Park, N. Arora, H. Huo, N. Maherali, T. Ahfeldt, A. Shimamura, M. W. Lensch, C. Cowan, K. Hochedlinger and G. Q. Daley, Cell, 2008, 134, 877–886 CrossRef CAS PubMed.
- J. Jang, J.-E. Yoo, J.-A. Lee, D. R. Lee, J. Y. Kim, Y. J. Huh, D.-S. Kim, C.-Y. Park, D.-Y. Hwang, H.-S. Kim, H.-C. Kang and D.-W. Kim, Exp. Mol. Med., 2012, 44, 202 CrossRef CAS PubMed.
- L. T. Ghaffari, A. Starr, A. T. Nelson and R. Sattler, Front. Neurosci., 2018, 12, 56 CrossRef PubMed.
- J. Lam, S. T. Carmichael, W. E. Lowry and T. Segura, Adv. Healthcare Mater., 2014, 4, 534–539 CrossRef PubMed.
- T. J. Rowland, L. M. Miller, A. J. Blaschke, E. L. Doss, A. J. Bonham, S. T. Hikita, L. V. Johnson and D. O. Clegg, Stem Cells Dev., 2010, 19, 1231–1240 CrossRef CAS PubMed.
- C. S. Hughes, L. M. Postovit and G. A. Lajoie, Proteomics, 2010, 10, 1886–1890 CrossRef CAS PubMed.
- M. Pakzad, M. Totonchi and A. Taei, Stem Cell Rev. Rep., 2010, 6, 96–107 CrossRef CAS PubMed.
- A. M. Kloxin, A. M. Kasko, C. N. Salinas and K. S. Anseth, Science, 2009, 324, 59–63 CrossRef CAS PubMed.
- A. M. Kloxin, J. A. Benton and K. S. Anseth, Biomaterials, 2010, 31, 1–8 CrossRef CAS PubMed.
- R. Y. Tam, L. J. Smith and M. S. Shoichet, Acc. Chem. Res., 2017, 50, 703–713 CrossRef CAS PubMed.
- S. A. Fisher, A. E. G. Baker and M. S. Shoichet, J. Am. Chem. Soc., 2017, 139, 7416–7427 CrossRef CAS PubMed.
- M. P. Lutolf, P. M. Gilbert and H. M. Blau, Nature, 2009, 462, 433–441 CrossRef CAS PubMed.
- J. A. Burdick and W. L. Murphy, Nat. Commun., 2012, 3, 1269 CrossRef PubMed.
- K. A. Mosiewicz, L. Kolb, A. J. van der Vlies, M. M. Martino, P. S. Lienemann, J. A. Hubbell, M. Ehrbar and M. P. Lutolf, Nat. Mater., 2013, 12, 1072–1078 CrossRef CAS PubMed.
- D. D. McKinnon, A. M. Kloxin and K. S. Anseth, Biomater. Sci., 2013, 1, 460–469 RSC.
- D. D. McKinnon, T. E. Brown, K. A. Kyburz, E. Kiyotake and K. S. Anseth, Biomacromolecules, 2014, 15, 2808–2816 CrossRef CAS PubMed.
- M. E. Smithmyer, L. A. Sawicki and A. M. Kloxin, Biomater. Sci., 2014, 2, 634–650 RSC.
- A. D. Celiz, J. G. W. Smith, R. Langer, D. G. Anderson, D. A. Winkler, D. A. Barrett, M. C. Davies, L. E. Young, C. Denning and M. R. Alexander, Nat. Mater., 2014, 13, 570–579 CrossRef CAS PubMed.
- M. R. Zanotelli, H. Ardalani, J. Zhang, Z. Hou, E. H. Nguyen, S. Swanson, B. K. Nguyen, J. Bolin, A. Elwell, L. L. Bischel, A. W. Xie, R. Stewart, D. J. Beebe, J. A. Thomson, M. P. Schwartz and W. L. Murphy, Acta Biomater., 2016, 35, 32–41 CrossRef CAS PubMed.
- S. Pellett, M. P. Schwartz, W. H. Tepp, R. Josephson, J. M. Scherf, C. L. Pier, J. A. Thomson, W. L. Murphy and E. A. Johnson, Sci. Rep., 2015, 5, 14566 CrossRef CAS PubMed.
- M. S. Rehmann, J. I. Luna, E. Maverakis and A. M. Kloxin, J. Biomed. Mater. Res., Part A, 2016, 104, 1162–1174 CrossRef CAS PubMed.
- S. T. Gould, N. J. Darling and K. S. Anseth, Acta Biomater., 2012, 8, 3201–3209 CrossRef CAS PubMed.
- A. M. Kloxin, M. W. Tibbitt and K. S. Anseth, Nat. Protoc., 2010, 5, 1867–1887 CrossRef CAS PubMed.
- C. M. Madl, B. L. Lesavage, R. E. Dewi, C. B. Dinh, R. S. Stowers, M. Khariton, K. J. Lampe, D. Nguyen, O. Chaudhuri, A. Enejder and S. C. Heilshorn, Nat. Mater., 2017, 16, 1233–1242 CrossRef CAS PubMed.
- X. Xu, Y. Tay, B. Sim, S. I. Yoon, Y. Huang, J. Ooi, K. H. Utami, A. Ziaei, B. Ng, C. Radulescu, D. Low, A. Y. J. Ng, M. Loh, B. Venkatesh, F. Ginhoux, G. J. Augustine and M. A. Pouladi, Stem Cell Rep., 2017, 8, 619–633 CrossRef CAS PubMed.
- E. Gabriel, A. Ramani, U. Karow, M. Gottardo, K. Natarajan, L. M. Gooi, G. Goranci-Buzhala, O. Krut, F. Peters, M. Nikolic, S. Kuivanen, E. Korhonen, T. Smura, O. Vapalahti, A. Papantonis, J. Schmidt-Chanasit, M. Riparbelli, G. Callaini, M. Krönke, O. Utermöhlen and J. Gopalakrishnan, Cell Stem Cell, 2017, 20, 397–406.e5 CrossRef CAS PubMed.
- B. D. Fairbanks, M. P. Schwartz, A. E. Halevi, C. R. Nuttelman, C. N. Bowman and K. S. Anseth, Adv. Mater., 2009, 21, 5005–5010 CrossRef CAS PubMed.
- J. Zhou, H. Y. Kim and L. A. Davidson, Development, 2009, 136, 677–688 CrossRef CAS PubMed.
- S. R. Caliari, M. Perepelyuk, B. D. Cosgrove, S. J. Tsai, G. Y. Lee, R. L. Mauck, R. G. Wells and J. A. Burdick, Sci. Rep., 2016, 6, 21387 CrossRef CAS PubMed.
- H. Shih, H.-Y. Liu and C.-C. Lin, Biomater. Sci., 2017, 5, 589–599 RSC.
- L. A. Sawicki and A. M. Kloxin, Biomater. Sci., 2014, 2, 1612–1626 RSC.
- G. Chen, Z. Hou, D. R. Gulbranson and J. A. Thomson, Cell Stem Cell, 2010, 7, 240–248 CrossRef CAS PubMed.
- Y. Yamada, K. Hozumi and M. Nomizu, Chem. – Eur. J., 2011, 17, 10500–10508 CrossRef CAS.
- B. E. Vogel, S. J. Lee, A. Hildebrand, W. Craig, M. D. Pierschbacher, F. Wong-Staal and E. Ruoslahti, J. Cell Biol., 1993, 121, 461–468 CrossRef CAS PubMed.
- S. E. Ochsenhirt, E. Kokkoli, J. B. McCarthy and M. Tirrell, Biomaterials, 2006, 27, 3863–3874 CrossRef CAS PubMed.
- M. Nomizu, W. H. Kim, K. Yamamura, A. Utani, S. Y. Song, A. Otaka, P. P. Roller, H. K. Kleinman and Y. Yamada, J. Biol. Chem., 1995, 270, 20583–20590 CrossRef CAS PubMed.
- R. Mhanna, A. Kashyap, G. Palazzolo, Q. Vallmajo-Martin, J. Becher, S. Möller, M. Schnabelrauch and M. Zenobi-Wong, Tissue Eng., Part A, 2014, 20, 1454–1464 CrossRef CAS PubMed.
- D. Zhang, M. Pekkanen-Mattila, M. Shahsavani, A. Falk, A. I. Teixeira and A. Herland, Biomaterials, 2014, 35, 1420–1428 CrossRef CAS PubMed.
- K. J. Lampe, A. L. Antaris and S. C. Heilshorn, Acta Biomater., 2013, 9, 5590–5599 CrossRef CAS PubMed.
- Q. Smith, E. Stukalin, S. Kusuma, S. Gerecht and S. X. Sun, Sci. Rep., 2015, 5, 12617 CrossRef CAS PubMed.
- Y. Feng and M. Mrksich, Biochemistry, 2004, 43, 15811–15821 CrossRef CAS PubMed.
- S. L. Vega, M. Y. Kwon, K. H. Song, C. Wang, R. L. Mauck, L. Han and J. A. Burdick, Nat. Commun., 2018, 9, 614 CrossRef PubMed.
- T. T. Yu and M. S. Shoichet, Biomaterials, 2005, 26, 1507–1514 CrossRef CAS PubMed.
- L. M. Y. Yu, F. D. Miller and M. S. Shoichet, Biomaterials, 2010, 31, 6987–6999 CrossRef CAS PubMed.
Footnote |
| † Electronic supplementary information (ESI) available. See DOI: 10.1039/c8bm00099a |
| This journal is © The Royal Society of Chemistry 2018 |