Characterizing the conformational ensemble of PROTAC degraders in solutions via atomistic simulations
Shikshya
Bhusal
 and
Omar
Valsson
and
Omar
Valsson
 *
*
Department of Chemistry, University of North Texas, Denton, Texas 76203, USA. E-mail: omar.valsson@unt.edu
First published on 6th November 2025
Abstract
Targeted protein degradation has emerged as a promising strategy for developing novel therapeutics, particularly for “undruggable” disease-related proteins. One approach is the use of PROteolysis TArgeting Chimeras (PROTACs) degraders, which induce the formation of ternary complexes between the target protein and E3 ligase, leading to ubiquitination and degradation of the target protein. Understanding the conformational behavior of PROTACs in solutions and how it relates to their pharmacokinetic properties and membrane permeability is crucial for optimizing PROTAC design and efficacy. Due to the large size and flexibility of PROTACs and their chameleonic character, it is essential to understand their conformational ensembles, and how they depend on the environment. Here, we introduce a novel methodology for exploring PROTAC conformational behavior using atomistic simulations. We employ the enhanced sampling method parallel bias metadynamics, where we bias generic local collective variables, specifically all rotatable dihedral angles, thereby avoiding the considerable challenge of identifying suitable global collective variables for biasing. The methodology allows for obtaining free energy surfaces of global CVs via reweighting. We apply the method to the prototypical case of the MZ1 PROTAC degrader, which targets bromodomain-containing protein-4 (BRD4) for degradation via the von Hippel-Lindau (VHL) E3 ligase, and elucidate its conformational behavior in different solvents, allowing us to gain insights into the chameleonic property of MZ1. Our results confirm that MZ1 adopts distinct conformations depending on the solvent, exhibiting collapsed conformations in water and chloroform, and extended conformations in DMSO. Collapsed conformations in chloroform have been correlated to increased cell permeability. Thus, our results show that MZ1 takes on conformations suitable for membrane permeation in apolar environments. Our methodological framework is generally applicable to large flexible molecules like PROTACs, and the results demonstrate its efficiency, laying the groundwork for similar investigations for other PROTACs and other “beyond-rule-of-5” drug candidates. This work provides valuable insights into the design and optimization of PROTACs, ultimately contributing to developing novel therapeutics for “undruggable” proteins.
1 Introduction
Proteins are integral to the regulation of crucial cellular processes that directly impact our health and survival. When these proteins exhibit abnormalities in their structure or function, it can lead to a variety of diseases. As a result, proteins have consistently emerged as vital targets for drug development for a long period of time. This focus on proteins has encouraged the creation of targeted therapies, including small-molecule inhibitors and precision-oriented antibodies, to address diseases such as cancer and neurological disorders.1Targeted protein degradation is a promising approach in the realm of therapeutics, offering solutions for proteins that have long resisted conventional drug targeting.2 This innovative field is gaining attention because it allows us to manipulate proteins that were previously challenging to target using standard small-molecule drugs and considered “undruggable”.3 These difficulties arise from factors such as the complex shapes of some protein active sites, which make them hard to bind to, or the smooth surfaces of certain proteins, which provide few binding sites for traditional drugs.4 PROteolysis TArgeting Chimeras (PROTACs) and molecular glues have emerged as alternative therapeutic modalities to target proteins.5 They proficiently overcome the limitations of traditional small-molecule inhibitors by leveraging the ubiquitin-proteasome system to tag and degrade target proteins, rather than merely inhibiting their activity. This mechanism allows for the targeting of “undruggable” proteins with complex structures or limited binding sites, enhancing selectivity and reducing off-target effects.5,6
PROTACs are highly specific molecules that degrade unwanted or harmful proteins in cells by hijacking the ubiquitin-proteasome system of the cell.5,7 The unique mechanism of PROTACs has attracted considerable attention in the pharmaceutical industry and shows promise in addressing a wide range of conditions, including cancers, immune disorders, viral infections, and neurodegenerative diseases.8 PROTAC molecules consist of two ligand-binding warheads that are connected via an intermediate linker. One of the ligands binds to the protein of interest, while the other one binds to an E3 ligase protein, inducing proximity and creating a ternary complex with a stable protein–protein interaction between the two proteins.9 Examples of E3 ligases are von Hippel-Lindau (VHL), cereblon (CRBN), MDM2, KEAP1, and CHIP.10 The role of E3 ligase in the cell is to tag protein for degradation by adding ubiquitin to it. Thus, by inducing proximity, the PROTAC molecule facilitates that the E3 ligase will ubiquitinate the protein of interest, leading to subsequent degradation in the proteasome.9 The intermediate linker also plays a vital role in physicochemical properties and bioactivity of the PROTAC molecule. The linker architecture determines to what degree the two ligands interact with the protein of interest and E3 ligase, and thus the maximal activity of the PROTAC molecule.11,12 An important aspect is that the PROTAC molecule is recycled and can engage another protein pair. A formidable challenge preventing PROTACs from realizing their therapeutic potential is their lack of compliance with the well-established drug-like properties associated with oral drugs, namely Lipinski's “rule-of-5” (Ro5) and Veber's rules.13,14 Over the past decade, the field of PROTACs has rapidly evolved, with numerous PROTACs developed for various E3 ligases and protein ligands, enabling the degradation of diverse proteins of interest. Numerous PROTACs are already in the advanced clinical trials including vepdegestrant (ARV-471).15 These advancements in PROTACs have resulted in achievements such as overcoming drug resistance, targeting previously “undruggable” proteins, and improving drug selectivity and specificity.2
Computational methods, including AI-driven drug design, molecular docking, molecular dynamics (MD) simulations, and quantum mechanics/molecular mechanics modeling, are increasingly essential in PROTAC development, enabling the prediction of degradation activity, structural modeling of ternary complexes, ubiquitination site accessibility, and optimization of linker properties and pharmacokinetics.9,16–24 These approaches have led to advances such as dual-target PROTACs and machine learning models capable of forecasting degradation efficiency and suggesting molecular modifications to enhance performance and selectivity.19,25
PROTACs are large and structurally complex molecules, comprising two ligand-binding warheads connected by a flexible linker group.9 This complex architecture poses significant challenges for membrane permeability and cellular uptake, ultimately affecting their bioavailability and intracellular activity. However, the flexibility of the linker group enables PROTACs to adopt multiple conformations, which is crucial for their function. This ability of PROTACs to change conformations or behavior in different environments is referred to as chameleonicity.26 This ability allows molecular chameleons to display both high aqueous solubility and high cell permeability.27 In the context of PROTACs, chameleonicity can be exhibited in several ways, such as changes in solubility, stability, or conformations in different solvents and cellular membrane environments, or in the presence of their target proteins and E3 ligases. For instance, PROTACs may adopt different conformations when bound to their target proteins and E3 ligases, forming ternary complexes that are necessary for their protein degradation activity.28 Moreover, this property can also impact their physicochemical properties, such as lipophilicity and polarity, which are critical for their oral bioavailability and pharmacokinetics.29 As such, understanding and optimizing PROTACs’ chameleonicity is an essential aspect of their development as therapeutic agents.
As a result of the flexible nature of the molecule and chamelonicity, studying a single conformation of PROTAC is insufficient, and instead, a conformational ensemble approach is necessary to capture their dynamical behavior. This approach, also explored in previous studies,30,31 involves characterizing the conformational behavior of PROTACs to understand their chameleonic behavior in different environments, such as aqueous and nonpolar solvents. By doing so, researchers can gain insights into how PROTACs adapt to various environments and maintain their activity, which is essential for their development as therapeutic agents. To accurately characterize these molecules, it is necessary to study the conformational ensemble, which can be represented as a collection of structures weighted by their Boltzmann factors.30 This approach allows researchers to capture the dynamic behavior of PROTACs and understand how they adapt to different environments. Atomistic molecular dynamics (MD) simulations are a powerful tool for studying the conformational ensemble of PROTACs, but they generally require enhanced sampling techniques32,33 to accurately sample the phase space and efficiently explore the vast conformational space. By employing these techniques, researchers can, in principle, reconstruct the free energy surface of PROTACs and their conformational ensemble and identify the most stable conformations, ultimately gaining insights into their chameleonic behavior and its implications for their therapeutic activity.
Dimethyl sulfoxide (DMSO), chloroform, and water are commonly used solvents to study the conformational behavior of PROTACs, as they mimic different cellular environments.34 Due to their varying polarities, these solvents help reveal how PROTAC structures adapt in polar versus apolar environments. DMSO supports permeability studies and is compatible with cellular systems.35 Chloroform plays an important role in mimicking the apolar environment of the cell membrane as it has a dielectric constant (ε = 4.8) similar to that of a lipid bilayer (ε = 3.0).11,36,37 The propensity of PROTACs to adopt collapsed conformations in apolar environments, such as chloroform, correlates with increased cell membrane permeability.11 Water, being highly polar, is ideal for exploring hydrophilic interactions and biological activity in physiological conditions. Together, these solvents provide a comprehensive picture of PROTAC behavior across biologically relevant environments.
In this study, we concentrate on the prototypical case of the MZ1 PROTAC molecule, that has been extensively studied both experimentally18,39–44 and using simulations.18,41 This compound induces the degradation of bromodomain-containing protein 4 (BRD4) by forming a ternary complex in a positively cooperative manner with a VHL E3 ligase, see Fig. 1(a). MZ1 is “sandwiched” between the two proteins, inducing extensive new protein–protein and protein–ligand contacts of both hydrophobic and electrostatic nature. It is known that BRD4 is overexpressed in a variety of tumor cells;45 thus, BRD4 has been an important target protein for degradation. Structurally, MZ1 consists of a VHL warhead linked to the BRD4 warhead via a three-unit poly(ethylene glycol) (PEG) linker, as shown in Fig. 1(c). The VHL warhead plays a critical role in recognizing and recruiting the E3 VHL ligase, which mediates the ubiquitination and subsequent degradation of the target protein, making it a key component in the function of PROTACs like MZ1.18,46,47
 | ||
| Fig. 1 (a) Crystal structure of the BRD4–MZ1–VHL ternary complex (PDB entry 5T3538,39), with BRD4 (red) and VHL (blue) bound by the PROTAC MZ1. (b) Molecular structure of the MZ1 PROTAC molecule. (c) Detailed view of MZ1, highlighting the BRD4 warhead, VHL warhead, and linker region. (d) The MZ1 molecule, highlighting 29 rotatable local dihedral angles used as collective variables in the parallel bias metadynamics simulations. | ||
The main aim of this paper is to develop a simulation methodology that allows us to accurately and efficiently characterize the conformational ensemble of PROTACs in different solvents, including water, chloroform, and DMSO. To achieve this, we employ the enhanced sampling method called parallel bias metadynamics (PBMetaD).48 A crucial aspect of enhanced sampling methods is the selection of collective variables (CVs), which can be challenging but is critical for the success of the simulation.49 The selection of inappropriate CVs can result in poor convergence and inaccurate outcomes. In this study, we address this challenge by considering generic local CVs, specifically all rotatable dihedral angles. We apply this strategy to the PROTAC molecule MZ1, identifying a total of 29 dihedral angles within the MZ1 structure. One of the benefits of parallel bias metadynamics is that it allows for biasing a large number of CVs simultaneously.48 A similar procedure was recently applied to characterize the conformational ensemble of intrinsically disordered peptides in solution.50–52 Employing generic local CVs like dihedral angles is a key aspect of our work as it allows for developing an effective sampling strategy that is generally applicable to most all PROTACs, and other large flexible molecules, but avoids the challenging task of selecting appropriate global CVs for biasing. By using this approach, we can efficiently explore the conformational space of MZ1 in different solvents and gain insights into its chameleonic behavior. Furthermore, we employ a reweighting method32,53–55 to remove the effect of the PBMetaD bias potential and obtain the free energy surface (FES) as a function of global CVs, such as radius of gyration, distances between different moieties within MZ1, solvent accessible surface area (SASA), and polar surface area (PSA). This approach enables us to obtain a more comprehensive understanding of the conformational ensemble of MZ1, including its overall shape, compactness, and solvent interactions. By analyzing these global CVs, we can gain insights into how the conformational ensemble of MZ1 changes in response to different solvents, and how these changes affect its chameleonic behavior and potential for therapeutic applications. In particular, understanding the conformational behavior in chloroform is of great importance as this apolar solvent mimics the hydrophobic membrane environment,11,36,37 and it has been observed that a higher likelihood of adopting collapsed conformations in chloroform correlates with increased cell permeability.11
2 Methods
We take the molecular structure for MZ1 from the crystal structure of the VHL–MZ1–BRD4 ternary complex from Protein Data Base (PDB) entry 5T35,38,39 see Fig. 1. The force field for MZ1 is parameterized with the general AMBER force field (GAFF)56 (Version 1.81, May 2017). The topology and parameter files are prepared using AmberTools22.57,58 To assign partial atomic charges for MZ1, we employ the restrained electrostatic potential (RESP)59,60 charge model using an electrostatic potential obtained from Hartree–Fock (HF) calculations using the 6-31G* basis set. The HF calculations are performed using the Gaussian 16 quantum chemistry package.61 We convert the MZ1 topology files from AMBER format to GROMACS format using ACPYPE.62 We consider three different solvent setups for MZ1, water, dimethyl sulfoxide (DMSO) and chloroform. For the water simulations, we employ the TIP3P water model.63 For DMSO64 and chloroform,65 we take the molecular structures from PubChem and parameterize their force field using GAFF and HF/6-31G* RESP charges in the same manner as for MZ1. For all solvent setups, MZ1 is placed in a dodecahedron periodic boundary box with a buffer of 30 Å (3.0 nm) between the molecule and box edges to avoid self-interactions in extended conformations. The system is then solvated by adding the relevant solvent molecules, resulting in 12![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 523 solvent molecules for the water setup, 3070 solvent molecules for the DMSO setup, and 3808 solvent molecules for the chloroform setup. For the water solvent setup, we add a physiological solution of sodium and chlorine ions with a salt concentration of 0.15 molar. The ions parameters are taken from the Amber99sb force field.66–69
523 solvent molecules for the water setup, 3070 solvent molecules for the DMSO setup, and 3808 solvent molecules for the chloroform setup. For the water solvent setup, we add a physiological solution of sodium and chlorine ions with a salt concentration of 0.15 molar. The ions parameters are taken from the Amber99sb force field.66–69
All the molecular dynamics (MD) simulations are run using the GROMACS 2022.5 code70 patched with the PLUMED 2.8.2 enhanced sampling plug-in code.71,72 We use the leap-frog integrator with a 2 fs timestep, periodic boundary conditions with a dodecahedron box, Verlet cutoff scheme73 with a 10 Å (1 nm) cutoff distance for electrostatic and van der Waals interactions, and the particle mesh Ewald (PME) method for long-range electrostatics.74 We employ long-range dispersion corrections for the energy and pressure. All bonds between heavy atoms and hydrogens are constrained with LINCS algorithm.75 The rigid TIP3P water model is constrained using the SETTLE algorithm.76 To maintain a stable temperature and pressure during the simulations, we employ the stochastic velocity rescale thermostat77 with a coupling time constant of 0.1 ps and a reference temperature of 300 K, and isotropic stochastic cell rescaling barostat78 with a coupling time constant of 1.0 ps, reference pressure of 1 atm, and compressibility of 4.5 × 10−5 bar−1. To validate the compressibility parameter for different solvents, we calculated the densities of neat chloroform and DMSO and found them to be approximately similar to experimental values.
The initial setup of the molecule in each solvent is performed through a multi-step process. First, to remove bad contacts, we perform energy minimization of 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 steps using the steepest descent algorithm with default convergence criteria. This is followed by an NVT (constant number of particles, volume, and temperature) equilibration run at 300 K for 2 ns, and then an NPT (constant number of particles, pressure, and temperature) equilibration simulation at 300 K and 1 atm pressure for 2 ns, with both simulations using position restraints on the heavy atoms of the MZ1 molecule to allow the solvent to relax (force constant of 1000 kJ mol−1 nm−2). Finally, we perform an NPT equilibration without position restraints at 300 K and 1 atm pressure for 2 ns. The final frame from this run is used as the starting point for further production molecular dynamics simulations.
000 steps using the steepest descent algorithm with default convergence criteria. This is followed by an NVT (constant number of particles, volume, and temperature) equilibration run at 300 K for 2 ns, and then an NPT (constant number of particles, pressure, and temperature) equilibration simulation at 300 K and 1 atm pressure for 2 ns, with both simulations using position restraints on the heavy atoms of the MZ1 molecule to allow the solvent to relax (force constant of 1000 kJ mol−1 nm−2). Finally, we perform an NPT equilibration without position restraints at 300 K and 1 atm pressure for 2 ns. The final frame from this run is used as the starting point for further production molecular dynamics simulations.
The parallel bias metadynamics simulations48 are run for 5 μs in the NPT ensemble at 300 K and 1 atm pressure. As CVs for biasing, we consider all 29 rotatable local dihedral angles shown in Fig. 1(d). The fluctuations of the dihedral angles within the planar ring moieties of the ligands binding to BRD4 and VHL are largely determined by the rigid planar geometry rather than specific interactions or conformational transitions so we exclude them from the biasing, thereby minimizing the risk of introducing artificial correlations in the biasing and ensuring a more accurate description of the systems' conformational landscape. We update the PBMetaD bias potentials by depositing Gaussians every 0.5 ps. We employ a bias factor of 24 and an initial Gaussian height of 0.3 kJ mol−1. The Gaussian width for all the dihedral angle CVs is set to 0.2 radians. The bias potentials are expanded on grids ranging from −π and +π radians and using a default grid spacing of 1/5 of the CVs’ Gaussian width. For the water solvent setup, we perform three independent PBMetaD simulations, and for the DMSO and chloroform solvent setup, we perform two independent PBMetaD simulations, with each simulation starting using different random seeds for the thermostat and barostat.
We also perform PBMetaD simulations for the water solvent setup where, in addition to the 29 dihedral angle CVs, we consider the radius of gyration of the MZ1 molecule as an additional biased CV. The radius of gyration CV is defined using all the heavy atoms of MZ1. The Gaussian widths for the radius of gyration CV is set as 0.5 Å (0.05 nm). The bias potential for the radius of gyration is expanded on a grid with a range from 0.0 to 15 Å (0.0 to 1.5 nm) and using a default grid spacing of 1/5 of the CVs' Gaussian width. All other PBMetaD biasing parameters are the same as above. We perform two independent simulations for this computational setup.
For analysis, we save trajectory frames and CV values every 1 ps. For trajectory analysis, we use VMD 1.9.3.79 In addition to the local dihedral angles, we also consider several global CVs to analyze the simulations. These include the distances between the planar ring moieties, specifically the ring structure of MZ1 (5, 6, or 7 membered rings), selected from both the BRD4 and VHL warheads, as shown in Fig. S2 and Table S1 in the SI. We also consider the radius of gyration, defined using all the heavy atoms of MZ1. All these CVs are calculated using the PLUMED 2.8.2 enhanced sampling plug-in code.71,72 We also consider the solvent accessible surface area (SASA)80 and the polar surface area (PSA).81 For PSA, we defined it to include all polar atoms and atoms with high electronegativity and the potential to engage in polar interactions, specifically oxygen, nitrogen, sulfur, chlorine, and hydrogens attached to these atoms. As discussed below and in the SI, we also considered PSA defined using several other definitions. SASA and PSA are calculated using the Shrake and Rupley algorithm82 implemented in GROMACS 2022.5, employing a default probe radius of 1.4 Å (0.14 nm). We employ PLUMED to extract conformations from free energy minima by applying conditional filtering in the minima region based on ranges of CV values, as shown in Table S2 in the SI.
To obtain an unbiased estimate of the free energy surfaces, we employ reweighting to remove the effect of the PBMetaD bias potential.32,54,83 To calculate the weights, we employ last-bias reweighting where we load all the Gaussians deposited after 5 μs and calculate the statistical weight for each frame in the simulation trajectory assuming a fixed bias potential.83 Additionally, we perform last-bias reweighting at several intermediate time points like 100 ns, 200 ns, 500 ns, 700 ns, 1 μs, 2 μs, and 3 μs to assess the convergence of the simulations and the reweighting procedure as shown in Fig. S7–S9 in the SI. All densities and free energy surfaces are obtained using kernel density estimation in PLUMED 2.8.2 where we validate the bandwidths employed by comparing to one-dimensional free energy profiles obtained using discrete histograms. All density and free energy surface figures are made using matplotlib.84
The GROMACS and PLUMED input files required to reproduce the results reported in this paper are available on PLUMED-NEST (https://www.plumed-nest.org), the public repository of the PLUMED consortium,72 as plumID:25.006 at https://www.plumed-nest.org/eggs/25/006/.
3 Results
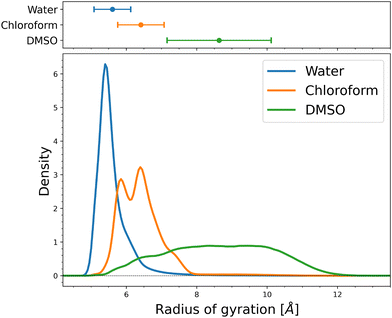
We perform the parallel bias metadynamics simulations for 5 μs at 300 K in three solvents: water, chloroform, and DMSO. While the local dihedral angles are useful as biased CVs in the parallel bias metadynamics simulations to obtain a comprehensive sampling of conformational space, on their own, they are generally not informative nor that useful in understanding the conformational behavior of the MZ1 molecule. Thus, we make use of the favorable property of metadynamics of being able to reweight the trajectory and obtain free energy surfaces as functions of any variables that are not biased. This allows us to characterize the conformational ensemble and how it differs between different environments by considering various global CVs.We start by considering the radius of gyration, which is a key global variable that provides valuable insights into the overall shape and compactness of a molecule. It measures the average distance of the atoms from the center of mass.85 Smaller radius of gyration values indicate more compact and globular structures, while larger values suggest more extended or unfolded conformations. Changes in the radius of gyration can signal conformational changes, such as folding or unfolding. Additionally, a compact molecule with a smaller radius of gyration may be less soluble in a solvent, whereas a more extended molecule with a larger radius of gyration may exhibit greater solubility.86 In Fig. 2, we present the distribution of the radius of gyration values obtained for the simulations in the different solvents. We observe that the radius of gyration values are quite low in water, as compared to the other solvent, indicating a more compact structure. This suggests that the molecule is more globular in water, potentially affecting its solubility and interactions with other molecules. In contrast, the radius of the gyration values are much larger in DMSO, indicating more extended and unfolded conformations. In addition, the radius of gyration distribution is quite broad in DMSO, indicating that the molecule can more easily sample different conformations, that is, it is more flexible, as compared to the water solvent. Similarly, like water, we observe that the radius of gyration values are also low for chloroform, indicating more collapsed structures than DMSO. Previous studies have shown that the tendency of PROTACs, such as MZ1, to adopt collapsed conformations in apolar environments like chloroform is associated with improved cell permeability.11 The distribution of radius of gyration for chloroform also shows a broader distribution than water but is not as broad as in DMSO. We can observe two peaks in the chloroform distribution that correspond to two distinct molecular conformations, as discussed below (see Fig. 6). These findings suggest that the molecule adopts different conformations in the three solvents, which may have implications for its behavior and function.
To understand the convergence behavior of our simulations, we obtain the radius of gyration distributions at different times and present the average and standard deviation as a function of time in Fig. 3. The corresponding radius of gyration distributions are shown in Fig. S7 in the SI. In Fig. 3, we can observe that the overall trends for the conformational behavior in the different solvents become evident after around 200–500 ns, while nearly full convergence is achieved after around 2 μs. The same conclusion can be obtained by considering Fig. S7 and S9 in the SI, where Fig. S9 exhibits the two-dimensional FES for the radius of gyration and distances between two warheads (see Fig. 4 below) obtained at different times. This demonstrates that shorter simulation times can be employed to characterize the differences in conformational behavior between solvents, especially when one is only interested in understanding trends, such as when characterizing and screening PROTACs in a drug discovery environment.
To ensure the reliability of our simulation procedure, we conducted multiple independent runs: three for water and two for both chloroform and DMSO. Analysis of global CVs revealed that all independent runs converged to the same results for each solvent, indicating strong consistency. The results for the water solvent setup are shown in Fig. S3–S5 in the SI, where further discussion on the methodological validation can be found. These results validate our methodology and demonstrate its effectiveness in sampling the conformational space of flexible PROTAC molecules like MZ1. We also perform a simulation for the water solvent setup where we consider the radius of gyration as an additional biased CV, in addition to the local dihedral angles CVs, similar to what was done in a previous study on an intrinsically disordered peptide.50 In these simulations, we obtain the same results for the global CVs as for the runs that only bias the dihedral angles as shown in Fig. S6 in the SI. This further validates our methodology and shows it is sufficient to consider only the local dihedral angles as biased CVs to obtain an accurate sampling of conformational space.
In addition to the radius of gyration, we consider various distances to illustrate the behavior of the MZ1 in different environments. Specifically, we identify important moieties in MZ1 from two warheads, as shown in Fig. S2 in the SI. We define the center of geometry of each moiety and calculate possible distances between these centers from both BRD4 and VHL warheads. One important distance that plays a significant role in our analysis is the end-to-end distance, or the distance between the two warheads, defined as the distance between the 6-membered ring moieties in the two warheads, as shown in Fig. 4(a). We show the distance distributions obtained in the different solvents in Fig. 4(b), where we can observe rather similar behavior as for the radius of gyration. The results indicate that in water, the molecule is compact and collapsed, while in DMSO it is more extended, flexible, and samples a wider range of conformations. The results in chloroform indicate that the conformations are more collapsed compared to those in DMSO. The collapsed conformations that we observed for MZ1 in apolar solvents like chloroform are significant, as it correlates with increased cell permeability.11 In Fig. 4(c), we present two-dimensional energy surfaces of the distance between the two warheads and the radius of gyration where we can observe that these two variables are highly correlated, as one might expect. The correlation is especially clear for DMSO. We also observe a single minimum in water, multiple minima in chloroform, and single broad minimum in DMSO.
Another important distance we consider is that between the tert-butyl group from the VHL warhead and the phenyl group from BRD4 warhead as shown in Fig. 5(a). From the results shown in Fig. 5(b) and (c), we can observe a similar pattern as before, where MZ1 is collapsed in water and more extended and flexible in DMSO.
To better understand the conformational behavior, we extract representative structures from the minima in the free energy surfaces for the distance between its two warheads and radius of gyration shown in Fig. 4(c). We illustrate the different MZ1 conformations in Fig. 6. As previously discussed, in water, the conformational landscape reveals a single minimum (panel a), corresponding to fully collapsed structures. In contrast, chloroform displays four distinct minima. The first minimum (panel b) resembles the collapsed conformations observed in water. The second (panel c) and third (panel d) minima also represent collapsed structures. However, the fourth minimum (panel e) shows semi-collapsed conformations. As discussed above, the chloroform radius of gyration distribution in Fig. 2 has two peaks. Here, the peak with radius of gyration values around 5.8 Å corresponds to structures with similar distance between warheads (panel c). The peak with radius of gyration values around 6.6 Å corresponds to structures with different distance between warheads (panels b, d, and e). For DMSO, the free energy minimum is much broader, so we analyze three representative areas. Structures obtained from the lower edge of the minimum (panel h) closely resemble the semi-collapsed conformation as seen in chloroform (panel e). Meanwhile, structures obtain from the center of the minimum (panel g) and the upper edge (panel f) show extended conformations of MZ1. This analysis provides a clearer understanding of how MZ1 adopts different conformations in water, chloroform, and DMSO.
 | ||
| Fig. 6 Representative MZ1 structures extracted from the minima of the free energy surfaces for the distance between its two warheads and radius of gyration (Fig. 4(c)). The blue box (panel (a)) shows conformations in water, the orange boxes (panels (b)–(e)) represent conformations in chloroform, and the green boxes (panels (f)–(h)) correspond to conformations in DMSO. These structures are obtained by extracting the conformations located within the small black box (indicating the low-free-energy region) shown in the figure. | ||
Solvent accessible surface area (SASA) is a key global collective variable that quantifies the total surface area of a biomolecule accessible to the solvent, see Fig. 7(a). This parameter provides valuable insights into the compactness and conformations of a molecule.80 Specifically, smaller SASA values indicates more compact, globular structures that have a smaller surface area exposed to the solvent, whereas larger SASA values suggests more extended or unfolded conformations with larger surface area accessible to the solvent. A molecule with larger SASA values may be more soluble in a solvent, as it has a larger surface area exposed to the solvent. Furthermore, SASA can be used to monitor important conformational changes in a molecule, such as folding or unfolding.87 Our results, depicted in Fig. 7, reveal a similar trend as above: SASA values are lower in water, indicating more compact structures, while they are higher in DMSO, suggesting extended conformations. Chloroform, on the other hand, exhibits SASA values lower than those in DMSO, reflecting collapsed conformations. We can observe from the two-dimensional free energy surfaces for SASA and the radius of gyration that these two global CVs are correlated, though the correlation is not as strong as for distances and the radius of gyration in Fig. 4 and 5.
Polar surface area (PSA) is a widely used molecular descriptor in drug transport studies, as it reflects the potential of a molecule to form hydrogen bonds and has been shown to associate with aqueous solubility and permeability.81 PSA is often correlated with the ability of a compound to pass through cell membranes and its potential for oral absorption, as molecules with high PSA typically exhibit reduced membrane permeability.88 In addition to its role in solubility and permeability, PSA can also be employed to estimate the potential for adverse drug reactions, such as interactions with transporters or enzymes in the body.14
There are different definitions of PSA used in literature; thus we have employed various definitions of PSA based on the existing literature to understand the differences among them. These different definitions are summarized in Table 1. First, we define PSA as the total surface area attributed solely to polar atoms and atoms with high electronegativity and the potential to engage in polar interactions, specifically oxygen, nitrogen, sulfur, chlorine.89 Second, PSA can be defined as the surface area contributed by selected atoms and the hydrogen atoms bonded to these electronegative atoms.81 Third, we define PSA as the combined surface area of selected atoms and carbon atoms that are either double-bonded to these electronegative atoms or bonded to two polar atoms. Finally, we expand the PSA definition to include the surface area from selected atoms, and hydrogen and carbon atoms included in the previous two definitions. It has been emphasized that incorporating carbon and hydrogen atoms in calculations such as PSA is essential, as these atoms significantly influence how a molecule interacts with its environment, including its solubility in solvents and its ability to bind to biological targets, which are critical factors in the design of effective drugs.14 The results for the different PSA definitions are shown in Fig. 8. We can observe that all definitions show a consistent trend of similar PSA values in water and chloroform, while PSA values in DMSO are higher. We can observe that including the hydrogens in PSA has a larger effect on differences between solvents (compare panels a and b), than including the attached carbon atoms (compare panels a and c). For PSA with only polar atoms (panel a), the water and chloroform values are almost the same. However, including the hydrogens in PSA (panel b) leads to a small relative shift of chloroform PSA to higher values compared to water. Adding carbon atoms (panel c) only leads to a near-constant shift of all distributions to higher values, due to PSA being calculated over larger number of atoms, while the relative differences between solvents are similar. These results indicate that it is beneficial to include the bonded hydrogen atoms in PSA calculation. Thus, in the following, we will employ the second definition of PSA calculated over the polar atoms and the hydrogen atoms bonded to these electronegative atoms, see Fig. 9(a).
| Definition | Selected atoms |
|---|---|
| 1 | Polar atoms and atoms with high electronegativity (N, S, O, Cl) |
| 2 | Polar atoms and atoms with high electronegativity + hydrogens attached to these atoms |
| 3 | Polar atoms and atoms with high electronegativity + carbons attached to two or more of these atoms |
| 4 | Polar atoms and atoms with high electronegativity + hydrogens attached to these atoms + carbons attached to two or more of these atoms |
 | ||
| Fig. 9 Conformational analysis of MZ1 in different solvents: (a) schematic representation of the MZ1 molecule, highlighting all the polar atoms and hydrogens attached to it, used to define polar surface area (PSA) (definition 2 in Table 1) (b) density plot of PSA in water, chloroform, and DMSO. The upper panel shows the average and standard deviation for each solvent. (c) Free energy surfaces of MZ1 as a function of PSA and the radius of gyration. | ||
Our results for PSA in different solvents in Fig. 9 reveal distinct solvation effects. Specifically, PSA in chloroform is lower than in the DMSO, reflecting the weaker interactions between the PROTAC molecule and the relatively non-polar environment of chloroform. In contrast, DMSO exhibits the highest PSA values (approximately 450 Å2), indicating stronger solute–solvent interactions due to the high polarity and hydrogen-bonding capacity of DMSO. For water and chloroform, the PSA value distributions have considerable overlap. In water, PSA values are slightly lower, with the polar atoms being less accessible to the solvent. In contrast, in chloroform, PSA values are slightly higher, indicating that the polar surface is more accessible to the solvent. These results suggest that the solvent's polarity and its ability to interact with the polar regions of the PROTAC molecule play a crucial role in determining the exposed polar surface area, which can influence the molecule's behavior in different environments. We also present two-dimensional free energy surfaces of PSA and the radius of gyration (panel c), where we can observe that the two variables are somewhat correlated, though the correlation is not as strong as for distances and the radius of gyration above (Fig. 4 and 5). Additionally, the results for other definitions of PSA are shown in SI, see Fig. S12–S14.
Interestingly, looking at the water and chloroform results, there is a slight difference between SASA and PSA values. SASA distinguishes the water and chloroform, though there is some overlap of the two distributions, with the chloroform SASA distribution tending to higher values than for water (see Fig. 7(b)). However, for PSA, the chloroform and water distributions have much more significant overlap, with the chloroform distribution tending to slightly higher values as compared to the water distribution for the second PSA definition (see Fig. 9(b)), though this depends on the definition of PSA employed as discussed above. The two-dimensional free energy surface of SASA and PSA is shown in Fig. S11 in the SI, where we can observe that these two variables are somewhat correlated, though the correlation appears to be less significant in water, as compared to chloroform and DMSO. These results indicate that SASA and PSA can measure different properties of the conformational ensemble.
4 Discussion
The solvent environment significantly influences the conformational behavior of the MZ1 molecule, as evidenced by analyzing key global variables such as the radius of gyration, SASA, and PSA. In water, the molecule adopts a more compact structure, as indicated by the reduced radius of gyration, SASA, and PSA values, showing that it is more globular. In addition, the water distributions are generally quite narrow, indicating a less flexible molecule. In DMSO, the molecule is more extended, with both a larger radius of gyration and SASA values. Furthermore, the DMSO distributions of the global variables are quite broad, indicating that the molecule can more easily sample different conformations and is more flexible. In chloroform, MZ1 also forms collapsed structures, but they are not as tightly collapsed as in water. These solvent-dependent conformational changes are further supported by the analysis of the end-to-end distances between different molecular moieties, revealing a similar trend in which the molecule is more collapsed in water and chloroform but more extended in DMSO. Additionally, PSA calculations show that the polar surface area of the PROTAC molecule is largest in DMSO, indicating stronger solute–solvent interactions due to DMSO's higher polarity and hydrogen-bonding capacity. In contrast, the PSA values are similar for water and chloroform, showing that the polar atoms are less exposed. Overall, the results underscore the crucial role of solvent polarity and solvation strength in modulating the structure and properties of drug-like molecules, which is vital for understanding their behavior in different chemical and biological environments.The conformational flexibility of the MZ1 molecule, influenced by the solvent environment, plays an important role in its bioavailability and membrane permeability. Since MZ1 is flexible, studying its conformational ensemble is essential, as a single structure does not adequately represent its behavior. In polar solvents like water, MZ1 may adopt more compact conformations, which can potentially facilitate higher membrane permeability by reducing their size and increasing lipophilicity. However, this compact form and the reduced flexibility can affect the molecule's ability to effectively engage both the target protein and the E3 ligase, which is crucial for its mechanism of action. Chloroform exhibits collapsed conformational behavior similar to that of water. Chloroform mimics the hydrophobic membrane environment, and it has been observed that the propensity to adopt collapsed conformations in chloroform correlates with increased cell permeability.11 Therefore, our observations of collapsed conformations in chloroform are significant, suggesting that in apolar environments, MZ1 takes on forms suitable for membrane permeation. This highlights the potential of chloroform to serve as a proxy for studying membrane interactions and permeability.
Our results are consistent with NMR experiments and MD simulation findings reported in a recent study on MZ1, which also observed similar conformational behavior under different solvent conditions, with MZ1 being collapsed in water and extended in DMSO.18
5 Conclusions
In conclusion, in this study, we introduce a novel methodological framework for exploring PROTAC conformational behavior using atomistic simulations, thereby, providing meaningful insights into the overall molecular behavior. The framework employs the enhanced sampling method parallel bias metadynamics to accurately explore the phase space by biasing local dihedral angles, followed by reweighting technique to compute free energy surfaces for global variables. By focusing on biasing local structural features rather than the complexity of identifying suitable global CVs for biasing, this approach simplifies the simulation process. This methodology is crucial for studying PROTACs, as these large and flexible molecules cannot be characterized by a single structure. Instead, their behavior and pharmacokinetic properties must be understood through their conformational ensemble. The application of this framework to the MZ1 PROTAC degrader in water, chloroform, and DMSO reveals distinct conformational behavior in each solvent: more compact in water and chloroform, and extended in DMSO. These conformational changes are reflected in differences in the radius of gyration, SASA, PSA, and inter-moiety distances, highlighting the adaptability of PROTACs in different environments and their chameleonic property. The conformational flexibility is key to the function of PROTACs and influences their membrane permeability. In apolar environments such as chloroform, collapsed conformations (as we observe for MZ1) correlate with increased cell permeability.11 Our results for MZ1 demonstrate the efficiency of this method in studying PROTACs, laying the groundwork for similar investigations for other PROTACs and other “beyond-rule-of-5” drug candidates,13 allowing valuable insights into the bioavailability and helping with drug design.Looking ahead, applying polarizable force fields could enhance the accuracy of the simulations by better capturing solvent effects and molecular interactions.90,91 Furthermore, machine learning potentials offer a promising direction for future research, as integrating such machine learning-based potentials with the current methodology could improve predictions of PROTAC behavior in diverse biological environments.92,93 Another key area for future work is the integration of experimental data (e.g., from NMR spectroscopy and small-angle X-ray scattering) with molecular simulations.94–97 This would refine and validate our models, ensuring that the simulated predictions closely align with real-world behavior.
Author contributions
Shikshya Bhusal: conceptualization, methodology, investigation, data curation, formal analysis, validation, visualization, writing – original draft, and writing – review and editing. Dr Omar Valsson: supervision, project administration, conceptualization, methodology, validation, writing – review and editing. All authors have read and agreed to the published version of the manuscript.Conflicts of interest
There are no conflicts of interest to declare.Data availability
Supplementary information with further details, results, and figures is available. See DOI: https://doi.org/10.1039/d5cp02530c.The GROMACS and PLUMED input files required to reproduce the results reported in this paper are available on PLUMED-NEST (https://www.plumed-nest.org), the public repository of the PLUMED consortium,72 as plumID:25.006 at https://www.plumed-nest.org/eggs/25/006/. Further data supporting the results reported in this paper is available upon request from the authors.
Acknowledgements
We thank Kresten Lindorff-Larsen (University of Copenhagen) and Simon Olsson (Chalmers University of Technology) for useful discussions. This work was supported by University of North Texas Start-up Funds provided to Dr Omar Valsson. All simulations were run on CRUNTCH4, the University of North Texas Chemistry Department HPC cluster that is supported by the National Science Foundation (NSF) under Award No. OAC-2117247.Notes and references
- A. Lai and C. Crews, Nat. Rev. Drug Discovery, 2017, 16, 101–114 CrossRef CAS PubMed.
- H. Pei, Y. Peng, Q. Zhao and Y. Chen, Small molecule PROTACs: An emerging technology for targeted therapy in drug discovery, 2019 Search PubMed.
- Q. Sun, H. Wang, J. Xie, L. Wang, J. Mu, J. Li, Y. Ren and L. Lai, Chem. Rev., 2025, 125, 6309–6365 CrossRef CAS PubMed.
- M. Békés, D. R. Langley and C. M. Crews, Nat. Rev. Drug Discovery, 2022, 21, 181–200 CrossRef PubMed.
- L. Zhao, J. Zhao and K. Zhong, et al. , Signal Transduction Targeted Ther., 2022, 7, 113 CrossRef CAS PubMed.
- Z. Liu, M. Hu and Y. Yang, et al. , Mol. Biomed., 2022, 3, 46 CrossRef CAS PubMed.
- K. M. Sakamoto, K. B. Kim, A. Kumagai, F. Mercurio, C. M. Crews and R. J. Deshaies, Proc. Natl. Acad. Sci. U. S. A., 2001, 98, 8554–8559 CrossRef CAS PubMed.
- M. He, C. Cao, Z. Ni, Y. Liu, P. Song, S. Hao, Y. He, X. Sun and Y. Rao, Signal Transduction Targeted Ther., 2022, 7, 181 CrossRef CAS PubMed.
- B. Mostofian, H.-J. Martin, A. Razavi, S. Patel, B. Allen, W. Sherman and J. A. Izaguirre, J. Chem. Inf. Model., 2023, 63, 5408–5432 CrossRef CAS PubMed.
- Y. Liu, J. Yang, T. Wang, M. Luo, Y. Chen, C. Chen, Z. Ronai, Y. Zhou, E. Ruppin and L. Han, Nat. Commun., 2023, 14, 6509 CrossRef CAS PubMed.
- V. Poongavanam, Y. Atilaw, S. Siegel, A. Giese, L. Lehmann, D. Meibom, M. Erdelyi and J. Kihlberg, J. Med. Chem., 2022, 65, 13029–13040 CrossRef CAS PubMed.
- V. Poongavanam, S. Peintner, Y. Abeje, F. Kölling, D. Meibom, M. Erdelyi and J. Kihlberg, ACS Med. Chem. Lett., 2025, 16, 681–687 CrossRef CAS PubMed.
- S. D. Edmondson, B. Yang and C. Fallan, Bioorg. Med. Chem. Lett., 2019, 29, 1555–1564 CrossRef CAS PubMed.
- D. F. Veber, S. R. Johnson, H.-Y. Cheng, B. R. Smith, K. W. Ward and K. D. Kopple, J. Med. Chem., 2002, 45, 2615–2623 CrossRef CAS PubMed.
- Arvinas and Pfizer, Arvinas and Pfizer's Vepde gestrant (ARV-471) Receives FDA Fast Track Designation for the Treatment of Patients with ER+/HER2-Metastatic Breast Cancer, 2024, https://ir.arvinas.com/news-releases/news-release-details/arvinas-and-pfizers-vepdegestrant-arv-471-receives-fda-fast, Accessed: 2025-01-27.
- M. L. Drummond, A. Henry, H. Li and C. I. Williams, J. Chem. Inf. Model., 2020, 60, 5234–5254 CrossRef CAS PubMed.
- X. Gao, H. A. Burris III, J. Vuky, R. Dreicer, A. O. Sartor, C. N. Sternberg, I. J. Percent, M. H. A. Hussain, A. Rezazadeh Kalebasty, J. Shen, E. I. Heath, G. Abesada-Terk, S. G. Gandhi, M. McKean, H. Lu, E. Berghorn, R. Gedrich, S. D. Chirnomas, N. J. Vogelzang and D. P. Petrylak, J. Clin. Oncol., 2022, 40, 17 Search PubMed.
- D. Weerakoon, R. J. Carbajo, L. D. Maria, C. Tyrchan and H. Zhao, J. Chem. Inf. Model., 2022, 62, 340–349 CrossRef CAS PubMed.
- F. Li, Q. Hu, X. Zhang, R. Sun, Z. Liu, S. Wu, S. Tian, X. Ma, Z. Dai, X. Yang, S. Gao and F. Bai, Nat. Commun., 2022, 13, 7133 CrossRef PubMed.
- G. Apprato, G. D’Agostini, P. Rossetti, G. Ermondi and G. Caron, Molecules, 2023, 28, 1206 CrossRef CAS PubMed.
- M. Ignatov, A. Jindal, S. Kotelnikov, D. Beglov, G. Posternak, X. Tang, P. Maisonneuve, G. Poda, R. A. Batey, F. Sicheri, A. Whitty, P. J. Tonge, S. Vajda and D. Kozakov, J. Am. Chem. Soc., 2023, 145, 7123–7135 CrossRef CAS PubMed.
- R. Tang, Z. Wang, S. Xiang, L. Wang, Y. Yu, Q. Wang, Q. Deng, T. Hou and H. Sun, JACS Au, 2023, 3, 1775–1789 CrossRef CAS PubMed.
- T. M. Tunjic, N. Weber and M. Brunsteiner, Comput. Struct. Biotechnol. J., 2023, 21, 2058–2067 CrossRef CAS PubMed.
- J. Ge, C.-Y. Hsieh, M. Fang, H. Sun and T. Hou, Trends Pharmacol. Sci., 2024, 45, 1162–1174 CrossRef CAS PubMed.
- D. Lv, P. Pal, X. Liu, Y. Jia, D. Thummuri, P. Zhang, W. Hu, J. Pei, Q. Zhang, S. Zhou, S. Khan, X. Zhang, N. Hua, Q. Yang, S. Arango, W. Zhang, D. Nayak, S. K. Olsen, S. T. Weintraub, R. Hromas, M. Konopleva, Y. Yuan, G. Zheng and D. Zhou, Nat. Commun., 2021, 12, 6896 CrossRef CAS PubMed.
- D. Lee, J. Choi, M. J. Yang, C.-J. Park and J. Seo, J. Med. Chem., 2023, 66, 13189–13204 CrossRef CAS PubMed.
- V. Poongavanam, L. Wieske and S. Peintner, et al. , Nat. Rev. Chem., 2024, 8, 45–60 CrossRef CAS PubMed.
- Y. Atilaw, V. Poongavanam, C. Svensson Nilsson, D. Nguyen, A. Giese, D. Meibom, M. Erdelyi and J. Kihlberg, ACS Med. Chem. Lett., 2021, 12, 107–114 CrossRef CAS PubMed.
- G. Apprato, V. Poongavanam, D. Garcia Jimenez, Y. Atilaw, M. Erdelyi, G. Ermondi, G. Caron and J. Kihlberg, Drug Discovery Today, 2024, 29, 103917 CrossRef CAS PubMed.
- M. Rossi Sebastiano, D. Garcia Jimenez, M. Vallaro, G. Caron and G. Ermondi, J. Med. Chem., 2022, 65, 12068–12083 CrossRef CAS PubMed.
- V. Poongavanam, et al. , J. Pharm. Sci., 2021, 110, 301–313 CrossRef CAS PubMed.
- J. Hénin, T. Lelièvre, M. R. Shirts, O. Valsson and L. Delemotte, Living J. Comput. Mol. Sci., 2022, 4, 1583 Search PubMed.
- S. Zheng and J. Pfaendtner, Mol. Simul., 2015, 41, 55–72 CrossRef CAS.
- G. Ermondi, D. G. Jimenez, M. Rossi Sebastiano, J. Kihlberg and G. Caron, Pharmaceutics, 2023, 15, 272 CrossRef CAS PubMed.
- A. Tjernberg, N. Markova, W. Griffiths and D. Hallén, J. Biomol. Screening, 2006, 11, 131–137 CrossRef CAS PubMed.
- G. Gramse, A. Dols-Perez, M. Edwards, L. Fumagalli and G. Gomila, Biophys. J., 2013, 104, 1257–1262 CrossRef CAS PubMed.
- Y. E. Abeje, L. H. E. Wieske, V. Poongavanam, S. Maassen, Y. Atilaw, P. Cromm, L. Lehmann, M. Erdelyi, D. Meibom and J. Kihlberg, J. Med. Chem., 2024, 68, 638–657 CrossRef PubMed.
- M. Gadd, M. Zengerle and A. Ciulli, The PROTAC MZ1 in complex with the second bromodomain of Brd4 and pVHL:ElonginC:ElonginB, 2017 DOI:10.2210/pdb5T35/pdb.
- M. S. Gadd, A. Testa, X. Lucas, K.-H. Chan, W. Chen, D. J. Lamont, M. Zengerle and A. Ciulli, Nat. Chem. Biol., 2017, 13, 514–521 CrossRef CAS PubMed.
- M. Zengerle, K. H. Chan and A. Ciulli, ACS Chem. Biol., 2015, 10, 1770–1777 CrossRef CAS PubMed.
- A. Testa, S. J. Hughes, X. Lucas, J. E. Wright and A. Ciulli, Angew. Chem., Int. Ed., 2019, 59, 1727–1734 CrossRef PubMed.
- F. J. Cimas, E. Niza, A. Juan, M. D. M. Noblejas-López, I. Bravo, A. Lara-Sanchez, C. Alonso-Moreno and A. Ocaña, Pharmaceutics, 2020, 12, 986 CrossRef CAS PubMed.
- G. Li, L. Ma, C. Feng, H. Yin, J. Bao, D. Wu, Z. Zhang, X. Li, Z. Li, C. Yang, H. Wang, F. Fang, X. Hu, M. Li, L. Xu, Y. Xu, H. Liang, T. Yang, J. Wang and J. Pan, BMC Cancer, 2024, 24, 220 CrossRef CAS PubMed.
- C. Crowe, M. A. Nakasone, S. Chandler, C. Craigon, G. Sathe, M. H. Tatham, N. Makukhin, R. T. Hay and A. Ciulli, Sci. Adv., 2024, 10, eado6492 CrossRef CAS PubMed.
- M. F. Segura, B. Fontanals-Cirera, A. Gaziel-Sovran, M. V. Guijarro, D. Hanniford, G. Zhang, P. González-Gomez, M. Morante, L. Jubierre, W. Zhang, F. Darvishian, M. Ohlmeyer, I. Osman, M.-M. Zhou and E. Hernando, Cancer Res., 2013, 73, 6264–6276 CrossRef CAS PubMed.
- M. L. Drummond and C. I. Williams, J. Chem. Inf. Model., 2019, 59, 1634–1644 CrossRef CAS PubMed.
- M. J. Roy, S. Winkler, S. J. Hughes, C. Whitworth, M. Galant, W. Farnaby, K. Rumpel and A. Ciulli, ACS Chem. Biol., 2019, 14, 361–368 CrossRef CAS PubMed.
- J. Pfaendtner and M. Bonomi, J. Chem. Theory Comput., 2015, 11, 5062–5067 CrossRef CAS PubMed.
- L. Bonati, V. Rizzi and M. Parrinello, J. Phys. Chem. Lett., 2020, 11, 2998–3004 CrossRef CAS PubMed.
- F. Pesce and K. Lindorff-Larsen, J. Phys. Chem. B, 2023, 127, 6277–6286 CrossRef CAS PubMed.
- T. Magsumov, I. Ibraev and I. Sedov, J. Phys. Chem. B, 2024, 128, 12333–12347 CrossRef CAS PubMed.
- R. K. Jain, C. K. Hall and E. E. Santiso, J. Phys. Chem. B, 2024, 128, 11164–11173 CrossRef CAS PubMed.
- M. Bonomi, A. Barducci and M. Parrinello, J. Comput. Chem., 2009, 30, 1615–1621 CrossRef CAS PubMed.
- P. Tiwary and M. Parrinello, J. Phys. Chem. B, 2015, 119, 736–742 CrossRef CAS PubMed.
- T. Shen and D. Hamelberg, J. Chem. Phys., 2008, 129, 034103 CrossRef PubMed.
- J. Wang, R. Wolf, J. Caldwell, P. Kollman and D. Case, J. Comput. Chem., 2004, 25, 1157–1174 CrossRef CAS PubMed.
- D. A. Case, T. E. Cheatham, T. Darden, H. Gohlke, R. Luo, K. M. Merz, A. Onufriev, C. Simmerling, B. Wang and R. J. Woods, J. Comput. Chem., 2005, 26, 1668–1688 CrossRef CAS PubMed.
- J. Wang, W. Wang, P. A. Kollman and D. A. Case, Antechamber, An Accessory Software Package For Molecular Mechanical Calculations, 2006, Department of Pharmaceutical Chemistry, University of California at San Francisco, San Francisco, California, 94143-0446 and Department of Molecular Biology, TPC15, The Scripps Research Institute, 10550 N. Torrey Pines Rd. La Jolla CA 92037.
- C. I. Bayly, P. Cieplak, W. Cornell and P. A. Kollman, J. Phys. Chem., 1993, 97, 10269–10280 CrossRef CAS.
- J. Wang, P. Cieplak and P. A. Kollman, J. Comput. Chem., 2000, 21, 1049–1074 CrossRef CAS.
- M. J. Frisch, G. W. Trucks, H. B. Schlegel, G. E. Scuseria, M. A. Robb, J. R. Cheeseman, G. Scalmani, V. Barone, G. A. Petersson, H. Nakatsuji, X. Li, M. Caricato, A. V. Marenich, J. Bloino, B. G. Janesko, R. Gomperts, B. Mennucci, H. P. Hratchian, J. V. Ortiz, A. F. Izmaylov, J. L. Sonnenberg, D. Williams-Young, F. Ding, F. Lipparini, F. Egidi, J. Goings, B. Peng, A. Petrone, T. Henderson, D. Ranasinghe, V. G. Zakrzewski, J. Gao, N. Rega, G. Zheng, W. Liang, M. Hada, M. Ehara, K. Toyota, R. Fukuda, J. Hasegawa, M. Ishida, T. Nakajima, Y. Honda, O. Kitao, H. Nakai, T. Vreven, K. Throssell, J. A. Montgomery, Jr., J. E. Peralta, F. Ogliaro, M. J. Bearpark, J. J. Heyd, E. N. Brothers, K. N. Kudin, V. N. Staroverov, T. A. Keith, R. Kobayashi, J. Normand, K. Raghavachari, A. P. Rendell, J. C. Burant, S. S. Iyengar, J. Tomasi, M. Cossi, J. M. Millam, M. Klene, C. Adamo, R. Cammi, J. W. Ochterski, R. L. Martin, K. Morokuma, O. Farkas, J. B. Foresman and D. J. Fox, Gaussian 16 Revision A.03, Gaussian Inc., Wallingford CT, 2016 Search PubMed.
- A. W. Sousa da Silva and W. F. Vranken, BMC Res. Notes, 2012, 5, 367 CrossRef PubMed.
- P. Mark and L. Nilsson, J. Phys. Chem. A, 2001, 105, 9954–9960 CrossRef CAS.
- N. C. for Biotechnology Information, PubChem Compound Summary for CID 679, Dimethyl Sulfoxide, 2024, https://pubchem.ncbi.nlm.nih.gov/compound/Dimethyl-Sulfoxide, Retrieved September 29, 2024.
- N. C. for Biotechnology Information, PubChem Compound Summary for CID 6212, Chloroform, 2024, https://pubchem.ncbi.nlm.nih.gov/compound/Chloroform, Retrieved September 29, 2024.
- K. Lindorff-Larsen, S. Piana, K. Palmo, P. Maragakis, J. L. Klepeis, R. O. Dror and D. E. Shaw, Proteins, 2010, 78, 1950–1958 CrossRef CAS PubMed.
- L. X. Dang, J. Am. Chem. Soc., 1995, 117, 6954–6960 CrossRef CAS.
- I. S. Joung and T. E. Cheatham, J. Phys. Chem. B, 2008, 112, 9020–9041 CrossRef CAS PubMed.
- J. Åqvist, J. Phys. Chem., 1990, 94, 8021–8024 CrossRef.
- M. J. Abraham, T. Murtola, R. Schulz, S. Páll, J. C. Smith, B. Hess and E. Lindahl, SoftwareX, 2015, 1–2, 19–25 CrossRef.
- G. A. Tribello, M. Bonomi, D. Branduardi, C. Camilloni and G. Bussi, Comput. Phys. Commun., 2014, 185, 604–613 CrossRef CAS.
- The PLUMED consortium, Nat. Methods, 2019, 16, 670–673 CrossRef PubMed.
- L. Verlet, Phys. Rev., 1967, 159, 98–103 CrossRef CAS.
- T. Darden, D. York and L. Pedersen, J. Chem. Phys., 1993, 98, 10089–10092 CrossRef CAS.
- B. Hess, J. Chem. Theory Comput., 2008, 4, 116–122 CrossRef CAS PubMed.
- S. Miyamoto and P. A. Kollman, J. Comput. Chem., 1992, 13, 952–962 CrossRef CAS.
- G. Bussi, D. Donadio and M. Parrinello, J. Chem. Phys., 2007, 126, 014101 CrossRef PubMed.
- M. Bernetti and G. Bussi, J. Chem. Phys., 2020, 153, 114107 CrossRef CAS PubMed.
- W. Humphrey, A. Dalke and K. Schulten, VMD: Visual Molecular Dynamics, 1996 Search PubMed.
- E. Durham, B. Dorr, N. Woetzel, R. Staritzbichler and J. Meiler, J. Mol. Model., 2009, 15, 1093–1108 CrossRef CAS PubMed.
- S. Prasanna and R. J. Doerksen, Curr. Med. Chem., 2009, 16, 21–41 CrossRef CAS PubMed.
- A. Shrake and J. Rupley, J. Mol. Biol., 1973, 79, 351–371 CrossRef CAS PubMed.
- D. Branduardi, G. Bussi and M. Parrinello, J. Chem. Theory Comput., 2012, 8, 2247–2254 CrossRef CAS PubMed.
- J. D. Hunter, Comput. Sci. Eng., 2007, 9, 90–95 Search PubMed.
- M. Y. Lobanov, N. S. Bogatyreva and O. V. Galzitskaya, Mol. Biol., 2008, 42, 623–628 CrossRef CAS.
- D. G. Amirabadi, Understanding the Radius of Gyration in Protein Design & Bioinformatics, 2024, Published Search PubMed.
- S. A. Ali, M. I. Hassan, A. Islam and F. Ahmad, Curr. Protein Pept. Sci., 2014, 15, 456–476 CrossRef CAS PubMed.
- P. Stenberg, K. Luthman and P. Artursson, J. Controlled Release, 2000, 65, 231–243 CrossRef CAS PubMed.
- P. Ertl, B. Rohde and P. Selzer, J. Med. Chem., 2000, 43, 3714–3717 CrossRef CAS PubMed.
- A. Warshel, M. Kato and A. V. Pisliakov, J. Chem. Theory Comput., 2007, 3, 2034–2045 CrossRef CAS PubMed.
- Z. Jing, C. Liu, S. Y. Cheng, R. Qi, B. D. Walker, J.-P. Piquemal and P. Ren, Annu. Rev. Biophys., 2019, 48, 371–394 CrossRef CAS PubMed.
- J. Vamathevan, D. Clark, P. Czodrowski, I. Dunham, E. Ferran, G. Lee, B. Li, A. Madabhushi, P. Shah, M. Spitzer and S. Zhao, Nat. Rev. Drug Discovery, 2019, 18, 463–477 CrossRef CAS PubMed.
- J. Jeon, S. Nim, J. Teyra, A. Datti, J. L. Wrana, S. S. Sidhu, J. Moffat and P. M. Kim, Genome Med., 2014, 6, 57 CrossRef PubMed.
- M. Bonomi, C. Camilloni, A. Cavalli and M. Vendruscolo, Sci. Adv., 2016, 2, e1501177 CrossRef PubMed.
- M. Bonomi, G. T. Heller, C. Camilloni and M. Vendruscolo, Curr. Opin. Struct. Biol., 2017, 42, 106–116 CrossRef CAS PubMed.
- M. Bonomi, C. Camilloni and M. Vendruscolo, Sci. Rep., 2016, 6, 31232 CrossRef CAS PubMed.
- J. Rappsilber, J. Struct. Biol., 2011, 173, 530–540 CrossRef CAS PubMed.
| This journal is © the Owner Societies 2025 |