Themed collection Computational Integrative biology (IB)

Front cover

Computational Integrative Biology – on the joint analysis of diverse biological data sets
Jan Baumbach and Richard Röttger introduce this Computational Integrative Biology themed issue.
Integr. Biol., 2014,6, 1008-1009
https://doi.org/10.1039/C4IB90037E
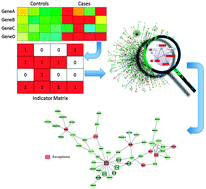
Modelling ligand selectivity of serine proteases using integrative proteochemometric approaches improves model performance and allows the multi-target dependent interpretation of features
Predicting ligand selectivity of serine proteases by integrating biological and chemical similarity into proteochemometric modelling approaches.
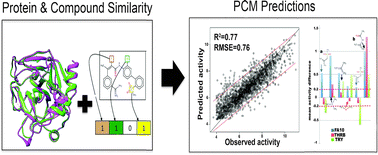
Integr. Biol., 2014,6, 1023-1033
https://doi.org/10.1039/C4IB00175C
C. pseudotuberculosis Phop confers virulence and may be targeted by natural compounds
The bacterial two-component system (TCS) regulates genes that are crucial for virulence in several pathogens.

Integr. Biol., 2014,6, 1088-1099
https://doi.org/10.1039/C4IB00140K
An improved interolog mapping-based computational prediction of protein–protein interactions with increased network coverage
Automated and efficient methods that map ortholog interactions from several organisms and public databases (pDB) are needed to identify new interactions in an organism of interest (interolog mapping).

Integr. Biol., 2014,6, 1080-1087
https://doi.org/10.1039/C4IB00136B
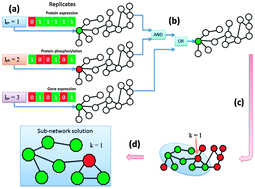
Integration of prior biological knowledge and epigenetic information enhances the prediction accuracy of the Bayesian Wnt pathway
Integration of prior biological knowledge and epigenetic information enhances the prediction accuracy of the Bayesian Wnt pathway making it a better diagnostic tool.
Integr. Biol., 2014,6, 1034-1048
https://doi.org/10.1039/C4IB00124A
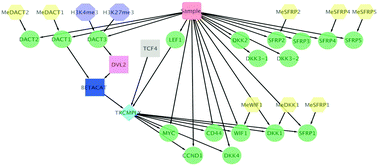
Elucidation of epithelial–mesenchymal transition-related pathways in a triple-negative breast cancer cell line model by multi-omics interactome analysis
Network features discriminate between epithelial and mesenchymal phenotype in a triple-negative breast cancer cell line model.
Integr. Biol., 2014,6, 1058-1068
https://doi.org/10.1039/C4IB00137K
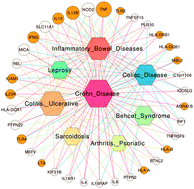
The integrated disease network
A disease network constructed by integrating biological data collected from diverse repositories is used for predicting novel disease–disease associations and drug repositioning opportunities.
Integr. Biol., 2014,6, 1069-1079
https://doi.org/10.1039/C4IB00122B
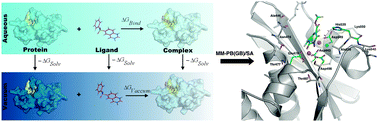
Binding free energy based structural dynamics analysis of HIV-1 RT RNase H–inhibitor complexes
The binding free energy based models have been used to study the structural dynamics of HIV-1 RT RNase H–inhibitor complexes.
Integr. Biol., 2014,6, 1010-1022
https://doi.org/10.1039/C4IB00111G
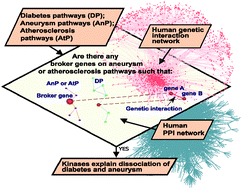
Network wiring of pleiotropic kinases yields insight into protective role of diabetes on aneurysm
Network wiring of pleiotropic kinases in disease pathways yields insight into protective role of diabetes in the development of aneurysm.
Integr. Biol., 2014,6, 1049-1057
https://doi.org/10.1039/C4IB00125G
About this collection
The discovery of differentiated new medicines that provide clear benefits to patients remains challenging in spite of ever increasing investments. At the same time the quantity and diversity of patient related data continues to grow exponentially (pre-clinical data, clinical data, patient data, EHR, ‘OMICs data, and information associated with medicines in general). The scientific community is starting to leverage this wealth of information in ways that have the potential to disrupt the traditional drug discovery and development process as we know it. This collection of articles was guest edited by Professor Jan Baumbach.