Small molecule induced poly(A) single strand to self-structure conformational switching: evidence for the prominent role of H-bonding interactions†
Abstract
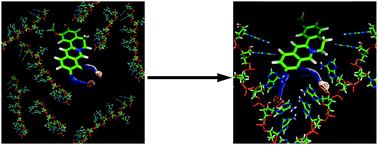
All messenger RNAs (mRNAs) have a polyadenylic acid tail that is added during post transcriptional RNA processing. Investigation of the structure–function and interactions of polyadenylic acid is an important area to target for cancer and related diseases. Jatrorrhizine and coptisine are two important isoquinoline alkaloids that are structurally very similar, differing only in the substituents on the isoquinoline chromophore. Here we demonstrate that these alkaloids differentially induce a self-structure in single stranded poly(A) using absorbance, thermal melting and differential scanning calorimetry experiments. Jatrorrhizine was found to be more effective than coptisine in binding to poly(A) from spectroscopy and calorimetry data. Molecular modeling results suggested the involvement of more H-bonds in the complexation of the former with poly(A). It appears that the presence of substituents on the alkaloid that can form H-bonding interactions with the adenine nucleotides may play a critical role in the binding and structural rearrangement of poly(A) into the self-structure. The atomic force microscopy data directly visualized the poly(A) self-structured network. We propose a plausible mechanism of the small molecule induced self-structure formation in poly(A). The results presented here may help in the design of effective poly(A) targeted molecules for therapeutic use.


 Please wait while we load your content...
Please wait while we load your content...