Bisulfite-free approaches for DNA methylation profiling
Abstract
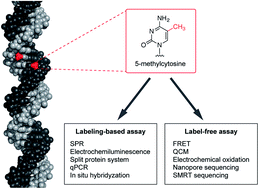
The determination of epigenetic modification, especially that of 5-methylcytosine in the CpG sequence in mammals, has attracted attention because it should prove valuable in a wide range of research fields including diagnosis, drug discovery and therapy. Various methods have been developed for recognizing 5-methylcytosine. Most of these methods employ bisulfite conversion, which mediates the deamination of unmethylated cytosine bases to uracil and leaves 5-methylcytosine. However, bisulfite conversion is time consuming with complex multiple operations and it leads to DNA degradation via its acid/base treatment. Therefore, bisulfite-free methods are required for DNA methylation analysis to allow epigenetic research to progress towards clinical applications. In this review, we introduce the recent development of bisulfite-free DNA methylation analysis, which we classify into two categories, namely labelling-based and labelling-free assays.


 Please wait while we load your content...
Please wait while we load your content...