Identification of key regulators and their controlling mechanism in a combinatorial apoptosis network: a systems biology approach†
Abstract
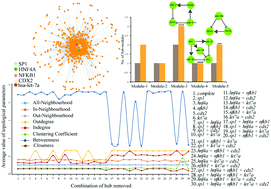
An experimentally validated set of apoptosis-regulatory proteins was subjected to network analysis, depicting a scale-free hierarchical fractal network. The power-law distribution of the various topological properties of the network revealed the fractal nature of the network, a signature of self-organization of the network where the network maintained the democratic constitution of nodes at various levels and showed the absence of the centrality–lethality control system. Even though network breakdown under the absence of the centrality–lethality rule of hub removal did not happen, the change in the topological properties of the network could be observed. Depending on the amount of change observed in topological properties, we identified a few proteins (hubs) which could be functionally important in the combinatorial apoptosis gene regulatory network. The crosstalk of these important proteins within the network along with functional modules probably tries to maintain the structural features of the network. NFKB1 was found to be the most efficient signal transducer followed by SP1. In addition, hsa-let-7a controlled the modules independently, revealing its importance in the incoherent and coherent types of feed forward loop motif analysis. The sub-modules and sub-sub-modules predicted bi-fan and multi-layer-perceptron motifs, suggesting the role of multifunctional signals in regulating apoptosis. Finally, SP1, NFKB1 and hsa-let-7a were observed to regulate apoptosis by influencing motifs, signal transduction, and module regulation, which was validated through the removal of hubs, signifying their biological importance in association with cancers.
 Please wait while we load your content...
Please wait while we load your content...