Significant differences in the biological activity of mononuclear Cu(ii) and Ni(ii) complexes with the polyquinolinyl ligand†
Abstract
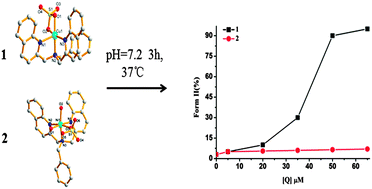
Two new mononuclear complexes [CuL(η2-SO4)] (1) and [NiL(η-SO4)(H2O)2]·3(H2O)·CH3CH2OH (2), where L is bis(2-quinolinyl methyl) benzyl-amine, have been synthesized and characterized by X-ray crystallography, ESI-MS and elemental analysis techniques. Complex 1 possesses a distorted square pyramidal geometry while complex 2 possesses distorted octahedral geometry. Interestingly, 2 dissolves in DMF–H2O mixed solvent, while 1 shows good water solubility. Both electronic absorption and EB displacement assay show that these complexes could bind to CT-DNA through partial intercalation. Though the DNA-binding abilities of 1 and 2 are similar, their chemical nuclease activities exhibit a great difference. Under physiological conditions, 1 could effectively convert pBR322 plasmid DNA from Form I to Form II at about 50 μM without any external agent, which shows self-oxidation cleavage activity; however, 2 shows poor DNA cleavage activity at 75 μM even in the presence of H2O2. At the same time, 1 also displays about 11-fold potential than 2 against HeLa cells with IC50 values 6.89 μM (1) and 74.85 μM (2). The IC50 value of 1 is even lower than the widely used drug cisplatin. The redox activity of Cu(II) and Ni(II) may play a main role in the biological activity of the two complexes, but the exact mechanism is still unclear. In addition, interactions of complexes with BSA have also been explored by fluorescence and UV-visible spectroscopic methods. The results indicated that complexes could quench the intrinsic fluorescence of BSA in a static quenching process.

 Please wait while we load your content...
Please wait while we load your content...