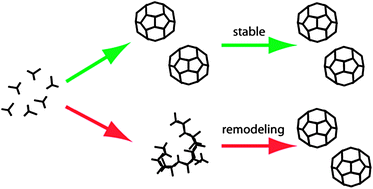
Dynamic remodelling of disordered protein aggregates is an alternative pathway to achieve robust self-assembly of nanostructures
Abstract
Clathrin is a naturally evolved
- This article is part of the themed collection: Directed self-assembly

 Please wait while we load your content...
Please wait while we load your content...