Uncovering the hidden geometry behind metabolic networks†
Abstract
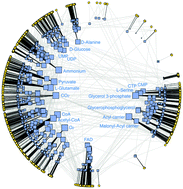
Metabolism is a fascinating cell machinery underlying life and disease and genome-scale reconstructions provide us with a captivating view of its complexity. However, deciphering the relationship between metabolic structure and function remains a major challenge. In particular, turning observed structural regularities into organizing principles underlying systemic functions is a crucial task that can be significantly addressed after endowing complex network representations of metabolism with the notion of geometric distance. Here, we design a cartographic map of metabolic networks by embedding them into a simple geometry that provides a natural explanation for their observed network topology and that codifies node proximity as a measure of hidden structural similarities. We assume a simple and general connectivity law that gives more probability of interaction to

 Please wait while we load your content...
Please wait while we load your content...