The in vivo fates of plant viral nanoparticles camouflaged using self-proteins: overcoming immune recognition
Abstract
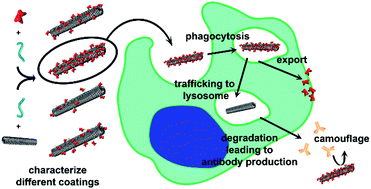
Nanoparticles offer a promising avenue for the targeted delivery of therapies. To slow clearance, nanoparticles are frequently stealth-coated to prevent opsonization and immune recognition. Serum albumin (SA) has been used as a bio-inspired stealth coating. To develop this shielding strategy for clinical applications, it is critical to understand the interactions between the immune system and SA-camouflaged nanoparticles. This work investigates the in vivo processing of SA-coated nanoparticles using tobacco mosaic virus (TMV) as a model system. In comparing four different SA-formulations, the particles with high SA coverage conjugated to TMV via a short linker performed the best at preventing antibody recognition. All formulations led to similar levels of TMV-specific antibodies after repeat administration in mice; importantly though, SA-specific antibodies were not detected and the TMV-specific antibodies were unable to recognize shielded SA-coated TMV. Upon uptake in macrophages, the shielding agent and nanoparticle separate, where TMV trafficked to the lysosome and SA appears to recycle. The distinct intracellular fates of the TMV carrier and SA shielding agent explain why anti-TMV but not SA-specific antibodies are generated. This work characterizes the outcomes of SA-camouflaged TMV after immune recognition, and highlights the effectiveness of SA as a nanoparticle shielding agent.


 Please wait while we load your content...
Please wait while we load your content...