Cells may feel a hard substrate even on a grafted layer of soft hydrogel†
Abstract
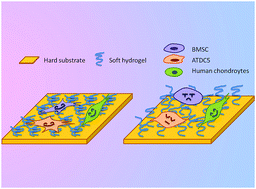
Introducing or grafting molecules onto biomaterial surfaces to regulate cell destination via biophysical cues is one of the important steps for biomaterial design in tissue engineering. Understanding how cells feel the substrate makes it easier to learn the mechanism behind cell–material interaction. In this study, on a glass substrate, we constructed poly-phenoxyethyl methacrylate (PHEMA) brushes having different lengths via a surface-induced atom transfer radical polymerization (SI-ATRP) method. FTIR-ATR and XPS tests of the formed polymer brushes indicate that these brushes have characteristic chemical structures of PHEMA; the polymer brush length revealed by the AFM tests increases linearly with reaction time. Cell lines of BMSCs, ATDC5, and human chondrocytes (HC) were cultured on these substrates to evaluate proliferation, adhesion, and differentiation. Our results demonstrated that the cells cultured on the substrates with short PHEMA brushes developed a spread morphology and organized actin fibers as compared to the cells cultured on those with long brushes. Different cell lines showed different responses depending on the PHEMA brush length. Cells cultured on long PHEMA brushes displayed a more rounded shape, higher gene expression of FAK and integrin, and lower gene expression of NCAM and N-cadherin as compared to those, especially ATDC5 cells, cultured on short PHEMA brushes. On PHEMA brushes with a long length, the cell lines express higher cartilage-specific genes including Sox9 and Col2 and GAG in ECM. The results suggest that polymer brushes having different lengths may interfere with the behavior of the cells cultured on them.


 Please wait while we load your content...
Please wait while we load your content...