Open Access Article
Open Access ArticleThrough tissue imaging of a live breast cancer tumour model using handheld surface enhanced spatially offset resonance Raman spectroscopy (SESORRS)†
Fay
Nicolson
 a,
Lauren E.
Jamieson
a,
Lauren E.
Jamieson
 a,
Samuel
Mabbott
a,
Samuel
Mabbott
 a,
Konstantinos
Plakas
a,
Konstantinos
Plakas
 b,
Neil C.
Shand
b,
Neil C.
Shand
 c,
Michael R.
Detty
c,
Michael R.
Detty
 b,
Duncan
Graham
b,
Duncan
Graham
 a and
Karen
Faulds
a and
Karen
Faulds
 *a
*a
aDepartment of Pure and Applied Chemistry, Technology and Innovation Centre, University of Strathclyde, 99 George Street, Glasgow G1 1RD, UK. E-mail: karen.faulds@strath.ac.uk
bDepartment of Chemistry, University at Buffalo, The State University of New York, New York 14260, USA
cDSTL, Porton Down, Salisbury, SP4 0JQ, UK
First published on 26th March 2018
Abstract
In order to improve patient survival and reduce the amount of unnecessary and traumatic biopsies, non-invasive detection of cancerous tumours is of imperative and urgent need. Multicellular tumour spheroids (MTS) can be used as an ex vivo cancer tumour model, to model in vivo nanoparticle (NP) uptake by the enhanced permeability and retention (EPR) effect. Surface enhanced spatially offset Raman spectroscopy (SESORS) combines both surface enhanced Raman spectroscopy (SERS) and spatially offset Raman spectroscopy (SORS) to yield enhanced Raman signals at much greater sub-surface levels. By utilizing a reporter that has an electronic transition in resonance with the laser frequency, surface enhanced resonance Raman scattering (SERRS) yields even greater enhancement in Raman signal. Using a handheld SORS spectrometer with back scattering optics, we demonstrate the detection of live breast cancer 3D MTS containing SERRS active NPs through 15 mm of porcine tissue. False color 2D heat intensity maps were used to determine tumour model location. In addition, we demonstrate the tracking of SERRS-active NPs through porcine tissue to depths of up to 25 mm. This unprecedented performance is due to the use of red-shifted chalcogenpyrylium-based Raman reporters to demonstrate the novel technique of surface enhanced spatially offset resonance Raman spectroscopy (SESORRS) for the first time. Our results demonstrate a significant step forward in the ability to detect vibrational fingerprints from a tumour model at depth through tissue. Such an approach offers significant promise for the translation of NPs into clinical applications for non-invasive disease diagnostics based on this new chemical principle of measurement.
Introduction
Non-invasive tumour detection is of vital importance for early cancer diagnosis, resulting in improved patient survival. Multicellular tumour spheroids (MTS) are used as tumour models and mimic in vivo tumour physiology. In contrast to traditional 2D monolayer cell cultures, these three dimensional cell models resemble the 3D in vivo environment more closely, establishing characteristic concentration gradients in oxygen, nutrients, and metabolites.1 MTS can be used as an ex vivo cancer model, avoiding the need for ethical approval and long term experiments associated with in vivo experiments. Termed the enhanced permeability and retention (EPR) effect, tumours retain nanoparticles (NPs) in their microenvironments due to poor lymphatic drainage. Thus, NP uptake is preferential in cancerous cells over healthy cell lines and NPs are frequently used to assist in imaging applications.2 MTS can be grown with a uniform distribution of NPs dispersed throughout and can provide a model for NP accumulation in tumours in vivo.3 MTS models were used here to develop a Raman imaging technique for effective tumour detection though tissue depths of several millimeters.Raman spectroscopy has previously been investigated for tumour detection,4 but is often limited to biopsy and cellular samples making it unable to non-invasively detect tumours at depth. Spatially offset Raman spectroscopy (SORS) provides a means of non-invasive investigation of tissue samples at depth by offsetting the point of collection from the point of laser excitation typically in the region of a few mm (Δx).5,6 Unlike confocal techniques where spectral information can be acquired at a particular depth by focusing the laser to a given distance in the z-axis, SORS makes use of a spatial offset to probe through the barrier/analyte system.5 SORS has shown promise in a number of applications including the detection of counterfeit alcohol,7 assessment of bone composition8,9 and in the analysis of cancerous calcifications in breast tissue.4 We have recently reported the use of a handheld SORS spectrometer for the detection of ethanol through up to 21 mm of plastic.10
When a molecule is adsorbed onto a metal NP surface, enhancement in scattering of the molecularly specific Raman reporter is observed. The surface enhanced Raman scattering (SERS) approach has been shown to yield enhancements several orders of magnitude greater than conventional Raman scattering.11 SERS has been applied in numerous applications including the sensitive detection of DNA,12 explosives detection,13 and in drug delivery sensing.14 Furthermore, through surface enhanced resonance Raman spectroscopy (SERRS), it is possible to greatly increase the Raman signal generated by the nanotags by using a laser that corresponds to an electronic transition of the Raman analyte to produce enhancements in the order of 1010 to 1014.15,16
Surface enhanced spatially offset Raman spectroscopy (SESORS) combines the benefits of SORS with SERS to achieve greater sample interrogation at depth.17 Using SESORS in a 180° back-scattering configuration, nanotags have been used in glucose sensing,18 tracked through 6.75 mm of tissue,19 and up to 8 mm of bone.20,21 Using SESORS in a transmission geometry, bisphosphonates have been tracked in bone through 20 mm of tissue22 and Stone et al. detected signal from nanotags through 45–50 mm of tissue.17 Several reports in the literature show the potential to target and image tumours in vivo using functionalized NPs.23,24 Kircher et al., functionalised Au nanoprobes with chalcogenpyrylium based dyes and the EGFR targeting antibody to successfully detect tumours in vivo using a mouse model.25 Whilst the authors reported attomolar limits of detection, little knowledge exists on the ability to detect tumours at clinically significant and relevant depths, and therefore the approach presented here provides a model to image and thus demonstrate, NP uptake in cancerous cells at notable depth. Here we report for the first time the use of NPs for the detection of a 3D breast tumour model buried at depths of 15 mm in tissue using a SORS instrument with back scattering optics. This was achieved by growing 3D breast cancer tumour models from cells incubated with SERRS active gold NPs (AuNPs) and then using the signal enhancing benefits of SERRS, combined with the through barrier detection capabilities of SORS, to locate the tumour model at depth using surface enhanced spatially offset resonance Raman spectroscopy (SESORRS) for the first time. In addition, these SESORRS measurements were carried out using a handheld spectrometer, which could potentially be used in a clinical environment. Furthermore, using the SESORRS approach, we report a penetration depth of up to 25 mm through tissue; the largest thickness to which SERS nanotags have been tracked using a back-scattering configuration.
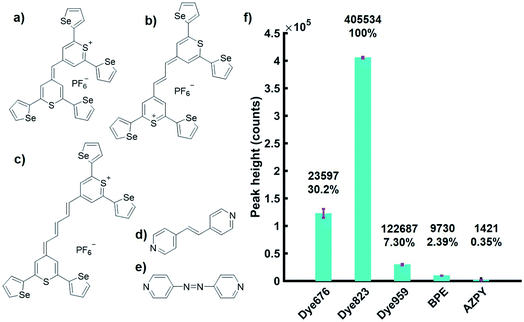
To achieve the detection of the tumour model at depth using the accumulation of NPs within a live 3D tumour model, we used chalcogenpyrylium dyes as resonant Raman reporters in combination with SESORRS detection. The absorbance wavelength of chalcogenpyrylium dyes can be fine-tuned into the near-infrared (NIR) to create a Raman reporter in resonance with the laser excitation wavelength of 830 nm.25 Our previous work has explored the use of red-shifted nanomaterials as SERS probes in the NIR by utilizing large AuNPs in conjunction with chalcogenpyrylium nanotags.26 Chalcogenpyrylium dyes conjugated to large AuNPs have been shown to generate strong SERS responses at picomolar detection levels at 1280 nm![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 27 and 1550 nm laser excitations.26 The chemical structure of each of the five Raman reporters used are shown in Fig. 1a–e. The absorbance wavelengths of the highly Raman active chalcogenpyrylium dyes; 676, 823, 959 (Fig. 1a–c), are tuned by increasing the number of sp2 carbons from 1 to 3 to 5 in the aliphatic backbone, thus, the wavelength of absorption is tuned to 676, 823 and 959 nm respectively.26,27 Further fine tuning is achieved through the choice of chalcogen atoms in the ring systems, which causes the absorption maximum to redshift. BPE (d) and AZPY (e) are non-resonant small molecules commonly used for SERS which were used for comparison. SERS spectra for each of these five Raman reporters are shown in the ESI Fig. S2.† All measurements were carried out using AuNPs with an average diameter of 83 nm. Each of the five Raman reporters were added to AuNPs to create nanotag solutions with a final dye concentration of 300 nM. The aim was to keep the dye concentration as low as possible by exploiting the benefit of using a Raman reporter that is in resonance with the laser. With regards to the experimental set up, all measurements were carried out using a total exposure time of 10 seconds (2 s integration time, 5 accumulations) at an 8 mm offset. The nose cone was fitted to use the instrument in a contact mode setting. The handheld instrument used here has a fixed maximum exposure time, therefore it is noted that if longer acquisition times were used, the signal to noise ratio may have improved.
27 and 1550 nm laser excitations.26 The chemical structure of each of the five Raman reporters used are shown in Fig. 1a–e. The absorbance wavelengths of the highly Raman active chalcogenpyrylium dyes; 676, 823, 959 (Fig. 1a–c), are tuned by increasing the number of sp2 carbons from 1 to 3 to 5 in the aliphatic backbone, thus, the wavelength of absorption is tuned to 676, 823 and 959 nm respectively.26,27 Further fine tuning is achieved through the choice of chalcogen atoms in the ring systems, which causes the absorption maximum to redshift. BPE (d) and AZPY (e) are non-resonant small molecules commonly used for SERS which were used for comparison. SERS spectra for each of these five Raman reporters are shown in the ESI Fig. S2.† All measurements were carried out using AuNPs with an average diameter of 83 nm. Each of the five Raman reporters were added to AuNPs to create nanotag solutions with a final dye concentration of 300 nM. The aim was to keep the dye concentration as low as possible by exploiting the benefit of using a Raman reporter that is in resonance with the laser. With regards to the experimental set up, all measurements were carried out using a total exposure time of 10 seconds (2 s integration time, 5 accumulations) at an 8 mm offset. The nose cone was fitted to use the instrument in a contact mode setting. The handheld instrument used here has a fixed maximum exposure time, therefore it is noted that if longer acquisition times were used, the signal to noise ratio may have improved.
To investigate the advantage of using resonant molecules for in vivo applications, nanotag solutions with each reporter were held in a quartz microcuvette. Porcine tissue samples of a 5 mm thickness were placed in front of the cuvette and brought into contact with the laser leaving no space between the nose cone and the sample (ESI, Fig. S3†). Spectra were acquired at an 8 mm spatial offset and truncated and baselined prior to processing. We have previously shown that at an 8 mm offset, the greatest level of through barrier detection takes place, which is the maximum capability of the instrument.10 The height of the most intense peak was then calculated as well as the relative percentage peak intensity (Fig. 1f). The handheld SORS instrument uses an excitation wavelength of 830 nm and it can be seen clearly in Fig. 1f that a significant enhancement in signal is generated by the resonant dye 823 compared to the off resonant molecules. Thus, by using a NIR resonant reporter molecule, superior SERRS signal through depth is generated over off resonant reporter molecules and the technique of SESORRS for greater through tissue detection is clearly demonstrated.
MTS were used as a 3D breast cancer tumour model to demonstrate the clinical significance of SESORRS for in vivo diagnostics. MCF7 human breast cancer cells were incubated overnight with the dye 823 nanotag solution (571 fM of AuNps, total of 7.092 × 1011 AuNp) resulting in uptake and accumulation of the nanotags within the cancer cells. MTS were then grown from a suspension of these cells using a hanging drop technique by pipetting 20 μL drops of MCF7 human breast cancer cell suspension onto the lid of a Petri dish. They were grown over a period of 9 days at 37 °C and 5% CO2 in a humidified incubator to a size <1 mm. No reduction in growth was observed. It is therefore reasonable to assume that the dyes did not cause cell death since cells need to be alive in order to divide and replicate to form MTS models. Previous reports show that NPs are homogenously dispersed throughout the MTS.3
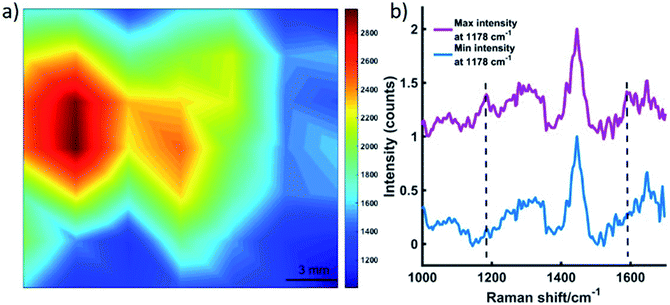
As described in Fig. S4, ESI,† roughly 10 MTS were transferred to a section of tissue (a). A 15 mm section of porcine tissue was then placed on top of the tissue layer upon which the MTS models were positioned (b). This model simulated the detection of SERRS active nanotags in tumours through tissue using the SORS approach. A translational x–y stage with a range of 2.54 cm was used to maneuver the tissue samples in steps of 3 mm to create an image of 7 × 7 pixels. As per the experimental set up described in Fig. S4, ESI,† the tissue system was brought into contact with the nose cone, and the z-value remained fixed, i.e. the stage was not moved in the z-direction. By fixing the z-direction, and utilising the SORS technique, we were able to probe through the tissue barrier to detect the MTS models through 15 mm of tissue. This was achieved by moving the stage the x–y direction to detect the MTS models through the tissue barrier. A false colour 2D SESORRS heat map of the peak intensity at 1178 cm−1 was then constructed, Fig. 2a. This corresponds to the uptake of dye 823 nanotags into MTS. Clear discrimination is seen between areas where the MTS were present and where they were not (Fig. 2a, b). There is a direct correlation between where the MTS models containing the SERRS-active nanotags were placed and the observed area of maximum intensity on the 2D map, i.e. the area of maximum intensity corresponds to the region in which the MTS models were positioned, (Fig. S4, ESI†). Similarly, at the point of minimum intensity in areas where the MTS models were not present, there was no spectral contribution from the dye (Fig. 2b). In this instance, the observed spectrum corresponds to that of the tissue. Therefore, since we know the precise location of the spheroids in the tissue, control spectra can be generated on the same image but away from the MTS deposition point. Furthermore, the SERRS-active NPs were contained within the spheroids themselves and therefore their location within in the system was known prior to imaging. The number of NPs present in each MTS was significantly less than for a bulk sample i.e. a nanotag solution held in a cuvette. However, despite this, the extremely sensitive detection of both dye 823 peaks at 1178 and 1592 cm−1 is confirmed in areas where the nanotags, and hence spheroid tumour models, were present (Fig. 2b), and thus the successful mapping of MTS through 15 mm of tissue using SESORRS was achieved. This proof of concept experiment demonstrates that targeted NPs in combination with SESORRS imaging could potentially be used to locate tumours at depth in vivo.
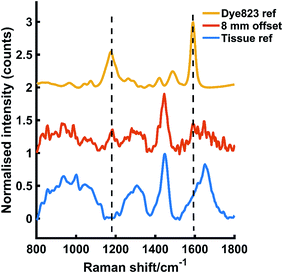
In a secondary, but no less significant demonstration of the power of this approach, dye 823 SERRS-active nanotags were detected through 25 mm of tissue. The nanotags were held in a quartz microcuvette (ESI Fig. S3†) and placed behind 25 mm of porcine tissue. Fig. 3 shows the dye 823 reference spectrum and tissue reference spectrum (top and bottom). The middle spectrum refers to SESORRS signal collected using an 8 mm offset. Both characteristic dye 823 peaks (1178 cm−1 and 1592 cm−1) are detectable by eye with the peak at 1178 cm−1 having greater distinction due to the lack of spectral overlap from the tissue in the 1600 cm−1 region. A scaled subtraction was also applied (Fig. S5, ESI†). This also confirms the detection of dye 823 demonstrating impressive potential of the SESORRS approach to detect nanotags through 25 mm of tissue using a back-scattering configuration by controlling the SERRS effect to achieve superior depth penetration. In previous collaborations with Stone and Matousek non-resonant SERS nanotags were detected through depths of up to 25 and 50 mm, however this was using a transmission geometry and a benchtop instrument.17 The results detailed here report the largest thickness that nanotags have been detected through using a back-scattering optical approach. Furthermore, all previous reports of SESORS has involved a benchtop system and this work highlights the ability of a handheld instrument to detect SESORRS signals through large thicknesses of tissue.
Through utilizing powerful chalcogenpyrylium-based Raman reporters for SERRS applications in combination with SORS we present the SESORRS technique for the first time, made possible through the resonant Raman tags, and report the highly significant and successful detection of 3D breast tumour models through 15 mm of tissue. To the best of our knowledge this is the first report of the detection of 3D tumour models using SESORRS. Furthermore, we show it is possible to detect nanotags through up to 25 mm of tissue, which is the largest thickness of through barrier detection reported using a back-scattering geometry, in contrast to a transmission approach. Previous work in the SESORS field has also involved benchtop systems, which are bulky and lack portability. The handheld instrument used in this instance has a fixed accumulation, i.e. the instrument permits only a certain maximum exposure time. Thus, whilst we would expect to achieve larger depth penetration and improved signal to noise with high-end benchtop systems which can facilitate longer acquisition times, particularly when resonant reporter molecules are used, the results described here show the excellent potential of handheld SORS which is more suited to clinical applications. Through the exploitation of the resonance effect, this novel work represents a significant step forward in the detection of vibrational fingerprints through tissue samples. Thus, an important step forward in the use of handheld back-scattering SESORRS for potential clinical applications including non-invasive tumour detection is demonstrated. Future work will focus on targeting and imaging tumours in vivo using SESORRS through the use of dual functionalised NPs, i.e. NPs functionalised with both a reporter molecule and a biomolecule that specifically targets the tumour in vivo. The use of a Raman reporter molecule will then facilitate the imaging of diseased tissue at depth due to accumulation of biofunctionalised NPs at the tumour site. Investigation of tumour depth location is also important and this will be investigated as the imaging capabilities of SORS develops to give a better insight into tumour location in vivo.
Funding
This work was supported by DSTL and the Engineering and Physical Sciences Research Council [grant numbers EP/J500550/1 and EP/M506643/1, KF and FN and EP/L014165/1, SM, DG and KF] and by the National Science Foundation [grant number CHE-1566142, KP and MRD]. Research data associated with this paper will become available through the following link: DOI: 10.15129/ef11d3c5-7c59-49b9-8009-143e52daa017.Conflicts of interest
The authors declare no conflict of interest.Acknowledgements
FN wishes to acknowledge the help of Stuart Bonthron at Cobalt Light Systems (now a part of Agilent).References
- L. E. Jamieson, D. J. Harrison and C. J. Campbell, Analyst, 2015, 140, 3910–3920 RSC.
- A. B. Chinen, C. M. Guan, J. R. Ferrer, S. N. Barnaby, T. J. Merkel and C. A. Mirkin, Chem. Rev., 2015, 115, 10530–10574 CrossRef CAS PubMed.
- L. E. Jamieson, V. L. Camus, P. O. Bagnaninchi, K. M. Fisher, G. D. Stewart, W. H. Nailon, D. B. McLaren, D. J. Harrison and C. J. Campbell, Nanoscale, 2016, 8, 16710–16718 RSC.
- N. Stone, R. Baker, K. Rogers, A. W. Parker and P. Matousek, Analyst, 2007, 132, 899–905 RSC.
- P. Matousek, Chem. Soc. Rev., 2007, 36, 1292–1304 RSC.
- P. Matousek and N. Stone, Chem. Soc. Rev., 2016, 45, 1794–1802 RSC.
- D. I. Ellis, R. Eccles, Y. Xu, J. Griffen, H. Muhamadali, P. Matousek, I. Goodall and R. Goodacre, Sci. Rep., 2017, 7, 12082 CrossRef PubMed.
- G. Feng, M. Ochoa, J. R. Maher, H. A. Awad and A. J. Berger, J. Biophotonics, 2017, 10, 990–996 CrossRef CAS PubMed.
- K. Sowoidnich, J. H. Churchwell, K. Buckely, A. E. Goodship, A. W. Parker and P. Matousek, Analyst, 2017, 142, 3219 RSC.
- F. Nicolson, L. E. Jamieson, S. Mabbott, N. C. Shand, D. Graham and K. Faulds, J. Raman Spectrosc., 2017, 48, 1828–1838 CrossRef CAS.
- S. Schlucker, Angew. Chem., Int. Ed., 2014, 53, 4756–4795 CrossRef PubMed.
- Y. C. Cao, R. Jin and C. A. Mirkin, Science, 2002, 297, 1536–1540 CrossRef CAS PubMed.
- S. S. R. Dasary, A. K. Singh, D. Senapati, H. Yu and P. C. Ray, J. Am. Chem. Soc., 2009, 131, 13806–13812 CrossRef CAS PubMed.
- J. Song, L. Pu, J. Zhou, B. Duan and H. Duan, ACS Nano, 2013, 7, 9947–9960 CrossRef CAS PubMed.
- B. Sharma, R. R. Frontiera, A.-I. Henry, E. Ringe and R. P. Van Duyne, Mater. Today, 2012, 15, 16–25 CrossRef CAS.
- S. McAughtrie, K. Faulds and D. Graham, J. Photochem. Photobiol., C, 2014, 21, 40–53 CrossRef CAS.
- N. Stone, M. Kerssens, G. R. Lloyd, K. Faulds, D. Graham and P. Matousek, Chem. Sci., 2011, 2, 776–780 RSC.
- K. Ma, J. M. Yuen, N. C. Shah, J. T. Walsh, M. R. Glucksberg and R. P. Van Duyne, Anal. Chem., 2011, 83, 9146–9152 CrossRef CAS PubMed.
- S. M. Asiala, N. C. Shand, K. Faulds and D. Graham, ACS Appl. Mater. Interfaces, 2017, 9, 25488–25494 CAS.
- B. Sharma, K. Ma, M. R. Glucksberg and R. P. Van Duyne, J. Am. Chem. Soc., 2013, 135, 17290–17923 CrossRef CAS PubMed.
- A. S. Moody, P. C. Baghernejad, K. R. Webb and B. Sharma, Anal. Chem., 2017, 89, 5688–5692 CrossRef CAS PubMed.
- H. N. Xie, R. Stevenson, N. Stone, A. Hernandez-Santana, K. Faulds and D. Graham, Angew. Chem., Int. Ed., 2012, 51, 8509–8511 CrossRef CAS PubMed.
- X. Qian, X.-H. Peng, D. O. Ansari, Q. Yin-Goen, G. Z. Chen, D. M. Shin, L. Yang, A. N. Young, M. D. Wang and S. Nie, Nat. Biotechnol., 2008, 26, 83–90 CrossRef CAS PubMed.
- A. Oseledchyk, C. Andreou, M. A. Wall and M. F. Kircher, ACS Nano, 2017, 11, 1488 CrossRef CAS PubMed.
- S. Harmsen, M. A. Bedics, M. A. Wall, R. Huang, M. R. Detty and M. F. Kircher, Nat. Commun., 2015, 6, 6570 CrossRef CAS PubMed.
- H. Kearns, M. A. Bedics, N. C. Shand, K. Faulds, M. R. Detty and D. Graham, Analyst, 2016, 141, 5062–5065 RSC.
- M. A. Bedics, H. Kearns, J. M. Cox, S. Mabbott, F. Ali, N. C. Shand, K. Faulds, J. B. Benedict, D. Graham and M. R. Detty, Chem. Sci., 2015, 6, 2302–2306 RSC.
Footnote |
| † Electronic supplementary information (ESI) available. See DOI: 10.1039/c8sc00994e |
| This journal is © The Royal Society of Chemistry 2018 |