Peptide-based nanoprobes for molecular imaging and disease diagnostics
Abstract
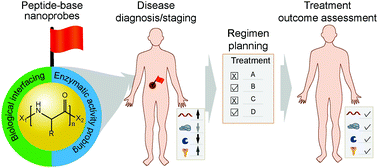
Pathological changes in a diseased site are often accompanied by abnormal activities of various biomolecules in and around the involved cells. Identifying the location and expression levels of these biomolecules could enable early-stage diagnosis of the related disease, the design of an appropriate treatment strategy, and the accurate assessment of the treatment outcomes. Over the past two decades, a great diversity of peptide-based nanoprobes (PBNs) have been developed, aiming to improve the in vitro and in vivo performances of water-soluble molecular probes through engineering of their primary chemical structures as well as the physicochemical properties of their resultant assemblies. In this review, we introduce strategies and approaches adopted for the identification of functional peptides in the context of molecular imaging and disease diagnostics, and then focus our discussion on the design and construction of PBNs capable of navigating through physiological barriers for targeted delivery and improved specificity and sensitivity in recognizing target biomolecules. We highlight the biological and structural roles that low-molecular-weight peptides play in PBN design and provide our perspectives on the future development of PBNs for clinical translation.
- This article is part of the themed collections: Peptide and protein nanotechnology and Probes for in vitro and in vivo fluorescence imaging


 Please wait while we load your content...
Please wait while we load your content...