Chemical strategies to unravel bacterial–eukaryotic signaling
Abstract
The common language of bacteria and higher life forms is a lexicon of small molecules that the research community is only beginning to decipher. While many new signaling molecules have been discovered in recent years, the identification of their targets is mostly lagging. This review will focus on the latest chemical-probe based research aimed at understanding how bacteria interact chemically with mammals and plants. In general, chemical biology strategies remain under-utilized in this complex field of research, with a few key exceptions, and we hope that this review encourages others to implement these techniques in their research. Specifically, we highlight the chemical biology techniques used in recent studies, especially activity-based protein profiling, that have been applied to unravel the chemical mechanisms of interkingdom interactions.
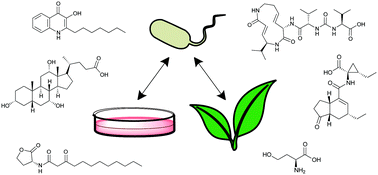
- This article is part of the themed collection: Chemical signaling at the eukaryotic/prokaryotic interface


 Please wait while we load your content...
Please wait while we load your content...