Defining the domains of Cia2 required for its essential function in vivo and in vitro†
Abstract
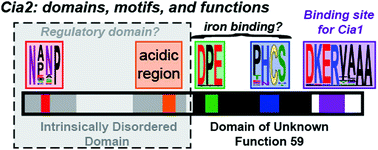
The cytosolic iron–sulfur cluster assembly (CIA) system biosynthesizes iron–sulfur (FeS) cluster cofactors for cytosolic and nuclear proteins. The yeast Cia2 protein is the central component of the targeting complex which identifies apo-protein targets in the final step of the pathway. Herein, we determine that Cia2 contains five conserved motifs distributed between an intrinsically disordered N-terminal domain and a C-terminal domain of unknown function 59 (DUF59). The disordered domain is dispensible for binding the other subunits of the targeting complex, Met18 and Cia1, and the apo-target Rad3 in vitro. While in vivo assays reveal that the C-terminal domain is sufficient to support viability, several phenotypic assays indicate that deletion of the N-terminal domain negatively impacts CIA function. We additionally establish that Glu208, located within a conserved motif found only in eukaryotic DUF59 proteins, is important for the Cia1–Cia2 interaction in vitro. In vivo, E208A–Cia2 results in a diminished activity of the cytosolic iron sulfur cluster protein, Leu1 but only modest effects on hydroxyurea or methylmethane sulfonate sensitivity. Finally, we demonstrate that neither of the two highly conserved motifs of the DUF59 domain are vital for any of Cia2's interactions in vitro yet mutation of the DPE motif in the DUF59 domain results in a nonfunctional allele in vivo. Our observation that four of the five highly conserved motifs of Cia2 are dispensable for targeting complex formation and apo-target binding suggests that Cia2 is not simply a protein–protein interaction mediator but it likely possesses an additional, currently cryptic, function during the final cluster insertion step of CIA.
- This article is part of the themed collection: Iron in Biology


 Please wait while we load your content...
Please wait while we load your content...