Receptor for hyaluronan mediated motility (RHAMM/HMMR) is a novel target for promoting subcutaneous adipogenesis
Abstract
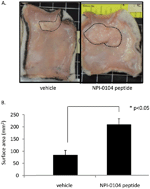
Hyaluronan, CD44 and the Receptor for Hyaluronan-Mediated Motility (RHAMM, gene name HMMR) regulate stem cell differentiation including mesenchymal progenitor differentiation. Here, we show that CD44 expression is required for subcutaneous adipogenesis, whereas RHAMM expression suppresses this process. We designed RHAMM function blocking peptides to promote subcutaneous adipogenesis as a clinical and tissue engineering tool. Adipogenic RHAMM peptides were identified by screening for their ability to promote adipogenesis in culture assays using rat bone marrow mesenchymal stem cells, mouse pre-adipocyte cell lines and primary human subcutaneous pre-adipocytes. Oil red O uptake into fat droplets and adiponectin production were used as biomarkers of adipogenesis. Positive peptides were formulated in either collagen I or hyaluronan (Orthovisc) gels then assessed for their adipogenic potential in vivo following injection into dorsal rat skin and mammary fat pads. Fat content was quantified and characterized using micro CT imaging, morphometry, histology, RT-PCR and ELISA analyses of adipogenic gene expression. Injection of screened peptides increased dorsal back subcutaneous fat pad area (208.3 ± 10.4 mm2versus control 84.11 ± 4.2 mm2; p < 0.05) and mammary fat pad size (45 ± 11 mg above control background, p = 0.002) in female rats. This effect lasted >5 weeks as detected by micro CT imaging and perilipin 1 mRNA expression. RHAMM expression suppresses while blocking peptides promote expression of PPARγ, C/EBP and their target genes. Blocking RHAMM function by peptide injection or topical application is a novel and minimally invasive method for potentially promoting subcutaneous adipogenesis in lipodystrophic diseases and a complementary tool to subcutaneous fat augmentation techniques.


 Please wait while we load your content...
Please wait while we load your content...