An electrochemical aptasensor for detection of bovine interferon gamma
Abstract
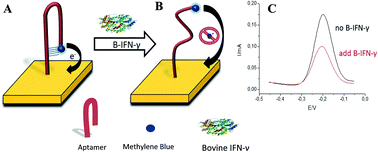
Bovine tuberculosis (TB) is an infectious disease caused by Mycobacterium tuberculosis (MTB) in which it is hard to identify the pathological symptoms. Release of bovine interferon gamma (BoIFN-γ) by T-cells provides an important diagnostic marker of MTB infections. In this work, we developed for the first time an electrochemical aptasensor for sensitive and specific determination of BoIFN-γ. A thiolated IFN-γ-binding aptamer was conjugated with methylene blue (MB) and immobilized on a gold electrode by self-assembly. Binding of IFN-γ to the electrode surface caused a conformational change in the aptamer, decreasing electron transfer efficiency. The redox current was quantified using square wave voltammetry (SWV) and was found to be specific for bovine IFN-γ with detection limit of 0.1 nM in pristine buffer and 0.9 nM in blood. The biosensor described here may, in the future, be used for on-site testing of bovine blood to help better identify and contain outbreaks of bovine TB.
- This article is part of the themed collection: RSC papers by NanoEngineering for Medicine and Biology 2018 Speakers


 Please wait while we load your content...
Please wait while we load your content...