Expanding the toolbox of organic chemists: directed evolution of P450 monooxygenases as catalysts in regio- and stereoselective oxidative hydroxylation
Abstract
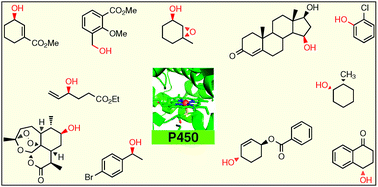
Cytochrome P450 enzymes (CYPs) have been used for more than six decades as catalysts for the CH-activating oxidative hydroxylation of organic compounds with formation of added-value products. However, it has not been possible to control regio- and stereoselectivity in a general manner, which is necessary for wide (industrial) applications. Especially directed evolution as a Darwinian approach to protein engineering has changed the situation for the better during the last 3–4 years leading to extensive progress, which is summarized and analysed in this Feature Article.
- This article is part of the themed collections: Celebrating the 150th anniversary of the German Chemical Society and Directed Evolution

 Please wait while we load your content...
Please wait while we load your content...