Nucleic acid therapeutics: basic concepts and recent developments
Abstract
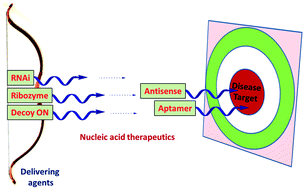
Nucleic acid based therapeutics have an allure for medicinal chemists due to their potential for specific control of gene expression. Persistent efforts have resulted in the FDA approval of the nucleic acid based drugs Vitravene™, Macugen™ and recently Kynamro™, leading to a sudden leap in the number of active clinical trials involving the nucleic acid moieties. Beginning with antigene and antisense technology, nucleic acid therapeutics have recently gained more potent strategies, e.g. RNA interference, ribozymes, decoy oligonucleotides and aptamers. Nucleic acid based therapeutics which have opened up new frontiers in medicinal chemistry research, along with various chemical modifications undertaken to improve their utility and their mechanism of action, will be discussed in this review article.
- This article is part of the themed collection: Chemistry for Medicine: Special Collection for RSC Advances

 Please wait while we load your content...
Please wait while we load your content...