Regulation of fatty acid and flavonoid biosynthesis by miRNAs in Lonicera japonica†
Abstract
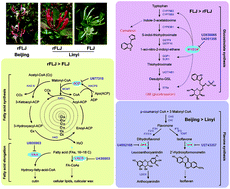
Lonicera japonica (honeysuckle) is an important herb with various pharmaceutically active secondary metabolites. MicroRNAs (miRNAs) are a class of small endogenous noncoding RNAs (sRNA), which play vital regulatory roles in plant secondary metabolism. Although sufficient data is available on the identification and functions of plant miRNAs, research on miRNA regulation of secondary metabolism in different varieties and producing areas of medical herbs has been scarce. In this study, we identified 28 conserved and 2517 novel miRNAs in honeysuckle using deep-sequencing analysis. The variety-specific regulation of miRNA expression was comparatively analysed by Cluster 3.0 algorithm. A total of 37 miRNAs showed differential expression among the three samples including two varieties of honeysuckle at two locations. The Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analyses indicated that 19 miRNAs with significantly different expression were putatively involved in the response against various factors and plant development. Additionally, the identification of target transcripts revealed that some of the miRNAs, namely U436803, U977315, U805963, U3938865, and U4351355, could be involved in fatty acid and secondary metabolite biosynthesis. To our knowledge, this is the first report on differentially expressed miRNAs in different varieties of honeysuckle. The results from this study can further facilitate the discovery of functional regulatory miRNAs involved in the secondary metabolism in L. japonica.


 Please wait while we load your content...
Please wait while we load your content...