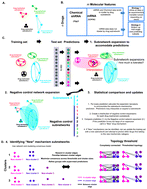
Molecular signatures are a powerful approach to characterize novel small molecules and derivatized small molecule libraries. While new experimental techniques are being developed in diverse model systems, informatics approaches lag behind these exciting advances. We propose an analysis pipeline for signature based drug annotation. We develop an integrated strategy, utilizing supervised and unsupervised learning methodologies that are bridged by network based statistics. Using this approach we can: 1, predict new examples of drug mechanisms that we trained our model upon; 2, identify “New” mechanisms of action that do not belong to drug categories that our model was trained upon; and 3, update our training sets with these “New” mechanisms and accurately predict entirely distinct examples from these new categories. Thus, not only does our strategy provide statistical generalization but it also offers biological generalization. Additionally, we show that our approach is applicable to diverse types of data, and that distinct biological mechanisms characterize its resolution of categories across different data types. As particular examples, we find that our predictive resolution of drug mechanisms from mRNA expression studies relies upon the analog measurement of a cell stress-related transcriptional rheostat along with a transcriptional representation of cell cycle state; whereas, in contrast, drug mechanism resolution from functional RNAi studies rely upon more dichotomous (e.g., either enhances or inhibits) association with cell death states. We believe that our approach can facilitate molecular signature-based drug mechanism understanding from different technology platforms and across diverse biological phenomena.
You have access to this article
 Please wait while we load your content...
Something went wrong. Try again?
Please wait while we load your content...
Something went wrong. Try again?


 Please wait while we load your content...
Please wait while we load your content...