From biological pathways to regulatory networks
Abstract
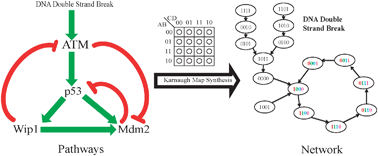
This paper presents a general theoretical framework for generating Boolean networks whose state transitions realize a set of given biological pathways or minor variations thereof. This ill-posed inverse problem, which is of crucial importance across practically all areas of biology, is solved by using Karnaugh maps which are classical tools for digital system design. It is shown that the incorporation of prior knowledge, presented in the form of biological pathways, can bring about a dramatic reduction in the cardinality of the network search space. Constraining the connectivity of the network, the number and relative importance of the attractors, and concordance with observed time-course data are additional factors that can be used to further reduce the cardinality of the search space. The networks produced by the approaches developed here should facilitate the understanding of multivariate biological phenomena and the subsequent design of intervention approaches that are more likely to be successful in practice. As an example, the results of this paper are applied to the widely studied p53 pathway and it is shown that the resulting network exhibits dynamic behavior consistent with experimental observations from the published literature.

 Please wait while we load your content...
Please wait while we load your content...