Refined kinetic transition networks for the GB1 hairpin peptide
Abstract
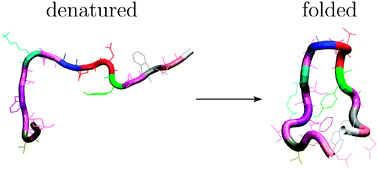
Refinement of databases of connected stationary points to describe global kinetics is discussed for the GB1 hairpin
Maintenance work is planned from 09:00 BST to 12:00 BST on Saturday 28th September 2024.
During this time the performance of our website may be affected - searches may run slowly, some pages may be temporarily unavailable, and you may be unable to access content. If this happens, please try refreshing your web browser or try waiting two to three minutes before trying again.
We apologise for any inconvenience this might cause and thank you for your patience.
* Corresponding authors
a
University Chemical Laboratories, Lensfield Road, Cambridge, UK
E-mail:
dw34@cam.ac.uk
Refinement of databases of connected stationary points to describe global kinetics is discussed for the GB1 hairpin

 Please wait while we load your content...
Something went wrong. Try again?
Please wait while we load your content...
Something went wrong. Try again?
J. M. Carr and D. J. Wales, Phys. Chem. Chem. Phys., 2009, 11, 3341 DOI: 10.1039/B820649J
To request permission to reproduce material from this article, please go to the Copyright Clearance Center request page.
If you are an author contributing to an RSC publication, you do not need to request permission provided correct acknowledgement is given.
If you are the author of this article, you do not need to request permission to reproduce figures and diagrams provided correct acknowledgement is given. If you want to reproduce the whole article in a third-party publication (excluding your thesis/dissertation for which permission is not required) please go to the Copyright Clearance Center request page.
Read more about how to correctly acknowledge RSC content.
 Fetching data from CrossRef.
Fetching data from CrossRef.
This may take some time to load.
Loading related content
